vedo.grids
Work with tetrahedral meshes.

1#!/usr/bin/env python3 2# -*- coding: utf-8 -*- 3import os 4import time 5from weakref import ref as weak_ref_to 6from typing import Any 7from typing_extensions import Self 8import numpy as np 9 10import vedo.vtkclasses as vtki # a wrapper for lazy imports 11 12import vedo 13from vedo import utils 14from vedo.core import PointAlgorithms 15from vedo.mesh import Mesh 16from vedo.file_io import download 17from vedo.visual import MeshVisual 18from vedo.transformations import LinearTransform 19 20__docformat__ = "google" 21 22__doc__ = """ 23Work with tetrahedral meshes. 24 25 26""" 27 28__all__ = [ 29 "UnstructuredGrid", 30 "TetMesh", 31 "RectilinearGrid", 32 "StructuredGrid", 33] 34 35 36######################################################################### 37class UnstructuredGrid(PointAlgorithms, MeshVisual): 38 """Support for UnstructuredGrid objects.""" 39 40 def __init__(self, inputobj=None): 41 """ 42 Support for UnstructuredGrid objects. 43 44 Arguments: 45 inputobj : (list, vtkUnstructuredGrid, str) 46 A list in the form `[points, cells, celltypes]`, 47 or a vtkUnstructuredGrid object, or a filename 48 49 Celltypes are identified by the following 50 [convention](https://vtk.org/doc/nightly/html/vtkCellType_8h_source.html). 51 """ 52 super().__init__() 53 54 self.dataset = None 55 56 self.mapper = vtki.new("PolyDataMapper") 57 self._actor = vtki.vtkActor() 58 self._actor.retrieve_object = weak_ref_to(self) 59 self._actor.SetMapper(self.mapper) 60 self.properties = self._actor.GetProperty() 61 62 self.transform = LinearTransform() 63 self.point_locator = None 64 self.cell_locator = None 65 self.line_locator = None 66 67 self.name = "UnstructuredGrid" 68 self.filename = "" 69 self.file_size = "" 70 71 self.info = {} 72 self.time = time.time() 73 self.rendered_at = set() 74 75 ###################################### 76 inputtype = str(type(inputobj)) 77 78 if inputobj is None: 79 self.dataset = vtki.vtkUnstructuredGrid() 80 81 elif utils.is_sequence(inputobj): 82 83 pts, cells, celltypes = inputobj 84 assert len(cells) == len(celltypes) 85 86 self.dataset = vtki.vtkUnstructuredGrid() 87 88 if not utils.is_sequence(cells[0]): 89 tets = [] 90 nf = cells[0] + 1 91 for i, cl in enumerate(cells): 92 if i in (nf, 0): 93 k = i + 1 94 nf = cl + k 95 cell = [cells[j + k] for j in range(cl)] 96 tets.append(cell) 97 cells = tets 98 99 # This would fill the points and use those to define orientation 100 vpts = utils.numpy2vtk(pts, dtype=np.float32) 101 points = vtki.vtkPoints() 102 points.SetData(vpts) 103 self.dataset.SetPoints(points) 104 105 # Fill cells 106 # https://vtk.org/doc/nightly/html/vtkCellType_8h_source.html 107 for i, ct in enumerate(celltypes): 108 if ct == vtki.cell_types["VERTEX"]: 109 cell = vtki.vtkVertex() 110 elif ct == vtki.cell_types["POLY_VERTEX"]: 111 cell = vtki.vtkPolyVertex() 112 elif ct == vtki.cell_types["TETRA"]: 113 cell = vtki.vtkTetra() 114 elif ct == vtki.cell_types["WEDGE"]: 115 cell = vtki.vtkWedge() 116 elif ct == vtki.cell_types["LINE"]: 117 cell = vtki.vtkLine() 118 elif ct == vtki.cell_types["POLY_LINE"]: 119 cell = vtki.vtkPolyLine() 120 elif ct == vtki.cell_types["TRIANGLE"]: 121 cell = vtki.vtkTriangle() 122 elif ct == vtki.cell_types["TRIANGLE_STRIP"]: 123 cell = vtki.vtkTriangleStrip() 124 elif ct == vtki.cell_types["POLYGON"]: 125 cell = vtki.vtkPolygon() 126 elif ct == vtki.cell_types["PIXEL"]: 127 cell = vtki.vtkPixel() 128 elif ct == vtki.cell_types["QUAD"]: 129 cell = vtki.vtkQuad() 130 elif ct == vtki.cell_types["VOXEL"]: 131 cell = vtki.vtkVoxel() 132 elif ct == vtki.cell_types["PYRAMID"]: 133 cell = vtki.vtkPyramid() 134 elif ct == vtki.cell_types["HEXAHEDRON"]: 135 cell = vtki.vtkHexahedron() 136 elif ct == vtki.cell_types["HEXAGONAL_PRISM"]: 137 cell = vtki.vtkHexagonalPrism() 138 elif ct == vtki.cell_types["PENTAGONAL_PRISM"]: 139 cell = vtki.vtkPentagonalPrism() 140 elif ct == vtki.cell_types["QUADRATIC_TETRA"]: 141 from vtkmodules.vtkCommonDataModel import vtkQuadraticTetra 142 cell = vtkQuadraticTetra() 143 elif ct == vtki.cell_types["QUADRATIC_HEXAHEDRON"]: 144 from vtkmodules.vtkCommonDataModel import vtkQuadraticHexahedron 145 cell = vtkQuadraticHexahedron() 146 elif ct == vtki.cell_types["QUADRATIC_WEDGE"]: 147 from vtkmodules.vtkCommonDataModel import vtkQuadraticWedge 148 cell = vtkQuadraticWedge() 149 elif ct == vtki.cell_types["QUADRATIC_PYRAMID"]: 150 from vtkmodules.vtkCommonDataModel import vtkQuadraticPyramid 151 cell = vtkQuadraticPyramid() 152 elif ct == vtki.cell_types["QUADRATIC_LINEAR_QUAD"]: 153 from vtkmodules.vtkCommonDataModel import vtkQuadraticLinearQuad 154 cell = vtkQuadraticLinearQuad() 155 elif ct == vtki.cell_types["QUADRATIC_LINEAR_WEDGE"]: 156 from vtkmodules.vtkCommonDataModel import vtkQuadraticLinearWedge 157 cell = vtkQuadraticLinearWedge() 158 elif ct == vtki.cell_types["BIQUADRATIC_QUADRATIC_WEDGE"]: 159 from vtkmodules.vtkCommonDataModel import vtkBiQuadraticQuadraticWedge 160 cell = vtkBiQuadraticQuadraticWedge() 161 elif ct == vtki.cell_types["BIQUADRATIC_QUADRATIC_HEXAHEDRON"]: 162 from vtkmodules.vtkCommonDataModel import vtkBiQuadraticQuadraticHexahedron 163 cell = vtkBiQuadraticQuadraticHexahedron() 164 elif ct == vtki.cell_types["BIQUADRATIC_TRIANGLE"]: 165 from vtkmodules.vtkCommonDataModel import vtkBiQuadraticTriangle 166 cell = vtkBiQuadraticTriangle() 167 elif ct == vtki.cell_types["CUBIC_LINE"]: 168 from vtkmodules.vtkCommonDataModel import vtkCubicLine 169 cell = vtkCubicLine() 170 elif ct == vtki.cell_types["CONVEX_POINT_SET"]: 171 from vtkmodules.vtkCommonDataModel import vtkConvexPointSet 172 cell = vtkConvexPointSet() 173 elif ct == vtki.cell_types["POLYHEDRON"]: 174 from vtkmodules.vtkCommonDataModel import vtkPolyhedron 175 cell = vtkPolyhedron() 176 elif ct == vtki.cell_types["HIGHER_ORDER_TRIANGLE"]: 177 from vtkmodules.vtkCommonDataModel import vtkHigherOrderTriangle 178 cell = vtkHigherOrderTriangle() 179 elif ct == vtki.cell_types["HIGHER_ORDER_QUAD"]: 180 from vtkmodules.vtkCommonDataModel import vtkHigherOrderQuadrilateral 181 cell = vtkHigherOrderQuadrilateral() 182 elif ct == vtki.cell_types["HIGHER_ORDER_TETRAHEDRON"]: 183 from vtkmodules.vtkCommonDataModel import vtkHigherOrderTetra 184 cell = vtkHigherOrderTetra() 185 elif ct == vtki.cell_types["HIGHER_ORDER_WEDGE"]: 186 from vtkmodules.vtkCommonDataModel import vtkHigherOrderWedge 187 cell = vtkHigherOrderWedge() 188 elif ct == vtki.cell_types["HIGHER_ORDER_HEXAHEDRON"]: 189 from vtkmodules.vtkCommonDataModel import vtkHigherOrderHexahedron 190 cell = vtkHigherOrderHexahedron() 191 elif ct == vtki.cell_types["LAGRANGE_CURVE"]: 192 from vtkmodules.vtkCommonDataModel import vtkLagrangeCurve 193 cell = vtkLagrangeCurve() 194 elif ct == vtki.cell_types["LAGRANGE_TRIANGLE"]: 195 from vtkmodules.vtkCommonDataModel import vtkLagrangeTriangle 196 cell = vtkLagrangeTriangle() 197 elif ct == vtki.cell_types["LAGRANGE_QUADRILATERAL"]: 198 from vtkmodules.vtkCommonDataModel import vtkLagrangeQuadrilateral 199 cell = vtkLagrangeQuadrilateral() 200 elif ct == vtki.cell_types["LAGRANGE_TETRAHEDRON"]: 201 from vtkmodules.vtkCommonDataModel import vtkLagrangeTetra 202 cell = vtkLagrangeTetra() 203 elif ct == vtki.cell_types["LAGRANGE_HEXAHEDRON"]: 204 from vtkmodules.vtkCommonDataModel import vtkLagrangeHexahedron 205 cell = vtkLagrangeHexahedron() 206 elif ct == vtki.cell_types["LAGRANGE_WEDGE"]: 207 from vtkmodules.vtkCommonDataModel import vtkLagrangeWedge 208 cell = vtkLagrangeWedge() 209 elif ct == vtki.cell_types["BEZIER_CURVE"]: 210 from vtkmodules.vtkCommonDataModel import vtkBezierCurve 211 cell = vtkBezierCurve() 212 elif ct == vtki.cell_types["BEZIER_TRIANGLE"]: 213 from vtkmodules.vtkCommonDataModel import vtkBezierTriangle 214 cell = vtkBezierTriangle() 215 elif ct == vtki.cell_types["BEZIER_QUADRILATERAL"]: 216 from vtkmodules.vtkCommonDataModel import vtkBezierQuadrilateral 217 cell = vtkBezierQuadrilateral() 218 elif ct == vtki.cell_types["BEZIER_TETRAHEDRON"]: 219 from vtkmodules.vtkCommonDataModel import vtkBezierTetra 220 cell = vtkBezierTetra() 221 elif ct == vtki.cell_types["BEZIER_HEXAHEDRON"]: 222 from vtkmodules.vtkCommonDataModel import vtkBezierHexahedron 223 cell = vtkBezierHexahedron() 224 elif ct == vtki.cell_types["BEZIER_WEDGE"]: 225 from vtkmodules.vtkCommonDataModel import vtkBezierWedge 226 cell = vtkBezierWedge() 227 else: 228 vedo.logger.error( 229 f"UnstructuredGrid: cell type {ct} not supported. Skip.") 230 continue 231 232 cpids = cell.GetPointIds() 233 cell_conn = cells[i] 234 for j, pid in enumerate(cell_conn): 235 cpids.SetId(j, pid) 236 self.dataset.InsertNextCell(ct, cpids) 237 238 elif "UnstructuredGrid" in inputtype: 239 self.dataset = inputobj 240 241 elif isinstance(inputobj, str): 242 if "https://" in inputobj: 243 inputobj = download(inputobj, verbose=False) 244 self.filename = inputobj 245 if inputobj.endswith(".vtu"): 246 reader = vtki.new("XMLUnstructuredGridReader") 247 else: 248 reader = vtki.new("UnstructuredGridReader") 249 self.filename = inputobj 250 reader.SetFileName(inputobj) 251 reader.Update() 252 self.dataset = reader.GetOutput() 253 254 else: 255 # this converts other types of vtk objects to UnstructuredGrid 256 apf = vtki.new("AppendFilter") 257 try: 258 apf.AddInputData(inputobj) 259 except TypeError: 260 apf.AddInputData(inputobj.dataset) 261 apf.Update() 262 self.dataset = apf.GetOutput() 263 264 self.properties.SetColor(0.89, 0.455, 0.671) # pink7 265 266 self.pipeline = utils.OperationNode( 267 self, comment=f"#cells {self.dataset.GetNumberOfCells()}", c="#4cc9f0" 268 ) 269 270 # ------------------------------------------------------------------ 271 def __str__(self): 272 """Print a string summary of the `UnstructuredGrid` object.""" 273 module = self.__class__.__module__ 274 name = self.__class__.__name__ 275 out = vedo.printc( 276 f"{module}.{name} at ({hex(self.memory_address())})".ljust(75), 277 c="m", bold=True, invert=True, return_string=True, 278 ) 279 out += "\x1b[0m\u001b[35m" 280 281 out += "nr. of verts".ljust(14) + ": " + str(self.npoints) + "\n" 282 out += "nr. of cells".ljust(14) + ": " + str(self.ncells) + "\n" 283 ct_arr = np.unique(self.cell_types_array) 284 cnames = [k for k, v in vtki.cell_types.items() if v in ct_arr] 285 out += "cell types".ljust(14) + ": " + str(cnames) + "\n" 286 287 if self.npoints: 288 out+="size".ljust(14)+ ": average=" + utils.precision(self.average_size(),6) 289 out+=", diagonal="+ utils.precision(self.diagonal_size(), 6)+ "\n" 290 out+="center of mass".ljust(14) + ": " + utils.precision(self.center_of_mass(),6)+"\n" 291 292 bnds = self.bounds() 293 bx1, bx2 = utils.precision(bnds[0], 3), utils.precision(bnds[1], 3) 294 by1, by2 = utils.precision(bnds[2], 3), utils.precision(bnds[3], 3) 295 bz1, bz2 = utils.precision(bnds[4], 3), utils.precision(bnds[5], 3) 296 out += "bounds".ljust(14) + ":" 297 out += " x=(" + bx1 + ", " + bx2 + ")," 298 out += " y=(" + by1 + ", " + by2 + ")," 299 out += " z=(" + bz1 + ", " + bz2 + ")\n" 300 301 for key in self.pointdata.keys(): 302 arr = self.pointdata[key] 303 rng = utils.precision(arr.min(), 3) + ", " + utils.precision(arr.max(), 3) 304 mark_active = "pointdata" 305 a_scalars = self.dataset.GetPointData().GetScalars() 306 a_vectors = self.dataset.GetPointData().GetVectors() 307 a_tensors = self.dataset.GetPointData().GetTensors() 308 if a_scalars and a_scalars.GetName() == key: 309 mark_active += " *" 310 elif a_vectors and a_vectors.GetName() == key: 311 mark_active += " **" 312 elif a_tensors and a_tensors.GetName() == key: 313 mark_active += " ***" 314 out += mark_active.ljust(14) + f': "{key}" ({arr.dtype}), ndim={arr.ndim}' 315 out += f", range=({rng})\n" 316 317 for key in self.celldata.keys(): 318 arr = self.celldata[key] 319 rng = utils.precision(arr.min(), 3) + ", " + utils.precision(arr.max(), 3) 320 mark_active = "celldata" 321 a_scalars = self.dataset.GetCellData().GetScalars() 322 a_vectors = self.dataset.GetCellData().GetVectors() 323 a_tensors = self.dataset.GetCellData().GetTensors() 324 if a_scalars and a_scalars.GetName() == key: 325 mark_active += " *" 326 elif a_vectors and a_vectors.GetName() == key: 327 mark_active += " **" 328 elif a_tensors and a_tensors.GetName() == key: 329 mark_active += " ***" 330 out += mark_active.ljust(14) + f': "{key}" ({arr.dtype}), ndim={arr.ndim}' 331 out += f", range=({rng})\n" 332 333 for key in self.metadata.keys(): 334 arr = self.metadata[key] 335 out += "metadata".ljust(14) + ": " + f'"{key}" ({len(arr)} values)\n' 336 337 return out.rstrip() + "\x1b[0m" 338 339 def _repr_html_(self): 340 """ 341 HTML representation of the UnstructuredGrid object for Jupyter Notebooks. 342 343 Returns: 344 HTML text with the image and some properties. 345 """ 346 import io 347 import base64 348 from PIL import Image 349 350 library_name = "vedo.grids.UnstructuredGrid" 351 help_url = "https://vedo.embl.es/docs/vedo/grids.html#UnstructuredGrid" 352 353 arr = self.thumbnail() 354 im = Image.fromarray(arr) 355 buffered = io.BytesIO() 356 im.save(buffered, format="PNG", quality=100) 357 encoded = base64.b64encode(buffered.getvalue()).decode("utf-8") 358 url = "data:image/png;base64," + encoded 359 image = f"<img src='{url}'></img>" 360 361 bounds = "<br/>".join( 362 [ 363 utils.precision(min_x, 4) + " ... " + utils.precision(max_x, 4) 364 for min_x, max_x in zip(self.bounds()[::2], self.bounds()[1::2]) 365 ] 366 ) 367 368 help_text = "" 369 if self.name: 370 help_text += f"<b> {self.name}:   </b>" 371 help_text += '<b><a href="' + help_url + '" target="_blank">' + library_name + "</a></b>" 372 if self.filename: 373 dots = "" 374 if len(self.filename) > 30: 375 dots = "..." 376 help_text += f"<br/><code><i>({dots}{self.filename[-30:]})</i></code>" 377 378 pdata = "" 379 if self.dataset.GetPointData().GetScalars(): 380 if self.dataset.GetPointData().GetScalars().GetName(): 381 name = self.dataset.GetPointData().GetScalars().GetName() 382 pdata = "<tr><td><b> point data array </b></td><td>" + name + "</td></tr>" 383 384 cdata = "" 385 if self.dataset.GetCellData().GetScalars(): 386 if self.dataset.GetCellData().GetScalars().GetName(): 387 name = self.dataset.GetCellData().GetScalars().GetName() 388 cdata = "<tr><td><b> cell data array </b></td><td>" + name + "</td></tr>" 389 390 pts = self.coordinates 391 cm = np.mean(pts, axis=0) 392 393 _all = [ 394 "<table>", 395 "<tr>", 396 "<td>", image, "</td>", 397 "<td style='text-align: center; vertical-align: center;'><br/>", help_text, 398 "<table>", 399 "<tr><td><b> bounds </b> <br/> (x/y/z) </td><td>" + str(bounds) + "</td></tr>", 400 "<tr><td><b> center of mass </b></td><td>" + utils.precision(cm,3) + "</td></tr>", 401 # "<tr><td><b> average size </b></td><td>" + str(average_size) + "</td></tr>", 402 "<tr><td><b> nr. points / cells </b></td><td>" 403 + str(self.npoints) + " / " + str(self.ncells) + "</td></tr>", 404 pdata, 405 cdata, 406 "</table>", 407 "</table>", 408 ] 409 return "\n".join(_all) 410 411 @property 412 def actor(self): 413 """Return the `vtkActor` of the object.""" 414 # print("building actor") 415 gf = vtki.new("GeometryFilter") 416 gf.SetInputData(self.dataset) 417 gf.Update() 418 out = gf.GetOutput() 419 self.mapper.SetInputData(out) 420 self.mapper.Modified() 421 return self._actor 422 423 @actor.setter 424 def actor(self, _): 425 pass 426 427 def _update(self, data, reset_locators=False): 428 self.dataset = data 429 if reset_locators: 430 self.cell_locator = None 431 self.point_locator = None 432 return self 433 434 def merge(self, *others) -> Self: 435 """ 436 Merge multiple datasets into one single `UnstrcturedGrid`. 437 """ 438 apf = vtki.new("AppendFilter") 439 for o in others: 440 if isinstance(o, UnstructuredGrid): 441 apf.AddInputData(o.dataset) 442 elif isinstance(o, vtki.vtkUnstructuredGrid): 443 apf.AddInputData(o) 444 else: 445 vedo.printc("Error: cannot merge type", type(o), c="r") 446 apf.Update() 447 self._update(apf.GetOutput()) 448 self.pipeline = utils.OperationNode( 449 "merge", parents=[self, *others], c="#9e2a2b" 450 ) 451 return self 452 453 def copy(self, deep=True) -> "UnstructuredGrid": 454 """Return a copy of the object. Alias of `clone()`.""" 455 return self.clone(deep=deep) 456 457 def clone(self, deep=True) -> "UnstructuredGrid": 458 """Clone the UnstructuredGrid object to yield an exact copy.""" 459 ug = vtki.vtkUnstructuredGrid() 460 if deep: 461 ug.DeepCopy(self.dataset) 462 else: 463 ug.ShallowCopy(self.dataset) 464 if isinstance(self, vedo.UnstructuredGrid): 465 cloned = vedo.UnstructuredGrid(ug) 466 else: 467 cloned = vedo.TetMesh(ug) 468 469 cloned.copy_properties_from(self) 470 471 cloned.pipeline = utils.OperationNode( 472 "clone", parents=[self], shape="diamond", c="#bbe1ed" 473 ) 474 return cloned 475 476 def bounds(self) -> np.ndarray: 477 """ 478 Get the object bounds. 479 Returns a list in format `[xmin,xmax, ymin,ymax, zmin,zmax]`. 480 """ 481 # OVERRIDE CommonAlgorithms.bounds() which is too slow 482 return np.array(self.dataset.GetBounds()) 483 484 def threshold(self, name=None, above=None, below=None, on="cells") -> Self: 485 """ 486 Threshold the tetrahedral mesh by a cell scalar value. 487 Reduce to only cells which satisfy the threshold limits. 488 489 - if `above = below` will only select cells with that specific value. 490 - if `above > below` selection range is flipped. 491 492 Set keyword "on" to either "cells" or "points". 493 """ 494 th = vtki.new("Threshold") 495 th.SetInputData(self.dataset) 496 497 if name is None: 498 if self.celldata.keys(): 499 name = self.celldata.keys()[0] 500 th.SetInputArrayToProcess(0, 0, 0, 1, name) 501 elif self.pointdata.keys(): 502 name = self.pointdata.keys()[0] 503 th.SetInputArrayToProcess(0, 0, 0, 0, name) 504 if name is None: 505 vedo.logger.warning("cannot find active array. Skip.") 506 return self 507 else: 508 if on.startswith("c"): 509 th.SetInputArrayToProcess(0, 0, 0, 1, name) 510 else: 511 th.SetInputArrayToProcess(0, 0, 0, 0, name) 512 513 if above is not None: 514 th.SetLowerThreshold(above) 515 516 if below is not None: 517 th.SetUpperThreshold(below) 518 519 th.Update() 520 return self._update(th.GetOutput()) 521 522 def isosurface(self, value=None, flying_edges=False) -> "vedo.mesh.Mesh": 523 """ 524 Return an `Mesh` isosurface extracted from the `Volume` object. 525 526 Set `value` as single float or list of values to draw the isosurface(s). 527 Use flying_edges for faster results (but sometimes can interfere with `smooth()`). 528 529 Examples: 530 - [isosurfaces1.py](https://github.com/marcomusy/vedo/tree/master/examples/volumetric/isosurfaces1.py) 531 532 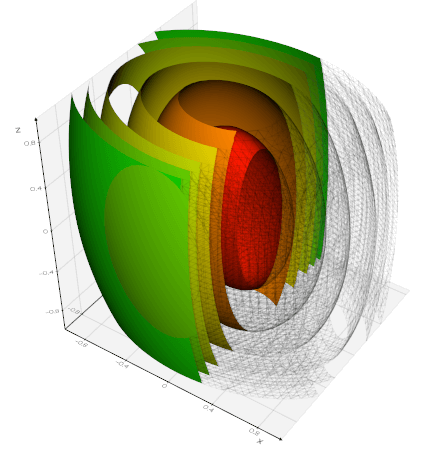 533 """ 534 scrange = self.dataset.GetScalarRange() 535 536 if flying_edges: 537 cf = vtki.new("FlyingEdges3D") 538 cf.InterpolateAttributesOn() 539 else: 540 cf = vtki.new("ContourFilter") 541 cf.UseScalarTreeOn() 542 543 cf.SetInputData(self.dataset) 544 cf.ComputeNormalsOn() 545 546 if utils.is_sequence(value): 547 cf.SetNumberOfContours(len(value)) 548 for i, t in enumerate(value): 549 cf.SetValue(i, t) 550 else: 551 if value is None: 552 value = (2 * scrange[0] + scrange[1]) / 3.0 553 # print("automatic isosurface value =", value) 554 cf.SetValue(0, value) 555 556 cf.Update() 557 poly = cf.GetOutput() 558 559 out = vedo.mesh.Mesh(poly, c=None).flat() 560 out.mapper.SetScalarRange(scrange[0], scrange[1]) 561 562 out.pipeline = utils.OperationNode( 563 "isosurface", 564 parents=[self], 565 comment=f"#pts {out.dataset.GetNumberOfPoints()}", 566 c="#4cc9f0:#e9c46a", 567 ) 568 return out 569 570 def shrink(self, fraction=0.8) -> Self: 571 """ 572 Shrink the individual cells. 573 574  575 """ 576 sf = vtki.new("ShrinkFilter") 577 sf.SetInputData(self.dataset) 578 sf.SetShrinkFactor(fraction) 579 sf.Update() 580 out = sf.GetOutput() 581 self._update(out) 582 self.pipeline = utils.OperationNode( 583 "shrink", comment=f"by {fraction}", parents=[self], c="#9e2a2b" 584 ) 585 return self 586 587 def tomesh(self, fill=False, shrink=1.0) -> "vedo.mesh.Mesh": 588 """ 589 Build a polygonal `Mesh` from the current object. 590 591 If `fill=True`, the interior faces of all the cells are created. 592 (setting a `shrink` value slightly smaller than the default 1.0 593 can avoid flickering due to internal adjacent faces). 594 595 If `fill=False`, only the boundary faces will be generated. 596 """ 597 gf = vtki.new("GeometryFilter") 598 if fill: 599 sf = vtki.new("ShrinkFilter") 600 sf.SetInputData(self.dataset) 601 sf.SetShrinkFactor(shrink) 602 sf.Update() 603 gf.SetInputData(sf.GetOutput()) 604 gf.Update() 605 poly = gf.GetOutput() 606 else: 607 gf.SetInputData(self.dataset) 608 gf.Update() 609 poly = gf.GetOutput() 610 611 msh = vedo.mesh.Mesh(poly) 612 msh.copy_properties_from(self) 613 msh.pipeline = utils.OperationNode( 614 "tomesh", parents=[self], comment=f"fill={fill}", c="#9e2a2b:#e9c46a" 615 ) 616 return msh 617 618 @property 619 def cell_types_array(self): 620 """Return the list of cell types in the dataset.""" 621 uarr = self.dataset.GetCellTypesArray() 622 return utils.vtk2numpy(uarr) 623 624 def extract_cells_by_type(self, ctype) -> "UnstructuredGrid": 625 """Extract a specific cell type and return a new `UnstructuredGrid`.""" 626 if isinstance(ctype, str): 627 try: 628 ctype = vtki.cell_types[ctype.upper()] 629 except KeyError: 630 vedo.logger.error(f"extract_cells_by_type: cell type {ctype} does not exist. Skip.") 631 return self 632 uarr = self.dataset.GetCellTypesArray() 633 ctarrtyp = np.where(utils.vtk2numpy(uarr) == ctype)[0] 634 uarrtyp = utils.numpy2vtk(ctarrtyp, deep=False, dtype="id") 635 selection_node = vtki.new("SelectionNode") 636 selection_node.SetFieldType(vtki.get_class("SelectionNode").CELL) 637 selection_node.SetContentType(vtki.get_class("SelectionNode").INDICES) 638 selection_node.SetSelectionList(uarrtyp) 639 selection = vtki.new("Selection") 640 selection.AddNode(selection_node) 641 es = vtki.new("ExtractSelection") 642 es.SetInputData(0, self.dataset) 643 es.SetInputData(1, selection) 644 es.Update() 645 646 ug = UnstructuredGrid(es.GetOutput()) 647 ug.pipeline = utils.OperationNode( 648 "extract_cell_type", comment=f"type {ctype}", c="#edabab", parents=[self] 649 ) 650 return ug 651 652 def extract_cells_by_id(self, idlist, use_point_ids=False) -> "UnstructuredGrid": 653 """Return a new `UnstructuredGrid` composed of the specified subset of indices.""" 654 selection_node = vtki.new("SelectionNode") 655 if use_point_ids: 656 selection_node.SetFieldType(vtki.get_class("SelectionNode").POINT) 657 contcells = vtki.get_class("SelectionNode").CONTAINING_CELLS() 658 selection_node.GetProperties().Set(contcells, 1) 659 else: 660 selection_node.SetFieldType(vtki.get_class("SelectionNode").CELL) 661 selection_node.SetContentType(vtki.get_class("SelectionNode").INDICES) 662 vidlist = utils.numpy2vtk(idlist, dtype="id") 663 selection_node.SetSelectionList(vidlist) 664 selection = vtki.new("Selection") 665 selection.AddNode(selection_node) 666 es = vtki.new("ExtractSelection") 667 es.SetInputData(0, self) 668 es.SetInputData(1, selection) 669 es.Update() 670 671 ug = UnstructuredGrid(es.GetOutput()) 672 pr = vtki.vtkProperty() 673 pr.DeepCopy(self.properties) 674 ug.actor.SetProperty(pr) 675 ug.properties = pr 676 677 ug.mapper.SetLookupTable(utils.ctf2lut(self)) 678 ug.pipeline = utils.OperationNode( 679 "extract_cells_by_id", 680 parents=[self], 681 comment=f"#cells {self.dataset.GetNumberOfCells()}", 682 c="#9e2a2b", 683 ) 684 return ug 685 686 def find_cell(self, p: list) -> int: 687 """Locate the cell that contains a point and return the cell ID.""" 688 if self.cell_locator is None: 689 self.cell_locator = vtki.new("CellLocator") 690 self.cell_locator.SetDataSet(self.dataset) 691 self.cell_locator.BuildLocator() 692 cid = self.cell_locator.FindCell(p) 693 return cid 694 695 def clean(self) -> Self: 696 """ 697 Cleanup unused points and empty cells 698 """ 699 cl = vtki.new("StaticCleanUnstructuredGrid") 700 cl.SetInputData(self.dataset) 701 cl.RemoveUnusedPointsOn() 702 cl.ProduceMergeMapOff() 703 cl.AveragePointDataOff() 704 cl.Update() 705 706 self._update(cl.GetOutput()) 707 self.pipeline = utils.OperationNode( 708 "clean", 709 parents=[self], 710 comment=f"#cells {self.dataset.GetNumberOfCells()}", 711 c="#9e2a2b", 712 ) 713 return self 714 715 def extract_cells_on_plane(self, origin: tuple, normal: tuple) -> Self: 716 """ 717 Extract cells that are lying of the specified surface. 718 """ 719 bf = vtki.new("3DLinearGridCrinkleExtractor") 720 bf.SetInputData(self.dataset) 721 bf.CopyPointDataOn() 722 bf.CopyCellDataOn() 723 bf.RemoveUnusedPointsOff() 724 725 plane = vtki.new("Plane") 726 plane.SetOrigin(origin) 727 plane.SetNormal(normal) 728 bf.SetImplicitFunction(plane) 729 bf.Update() 730 731 self._update(bf.GetOutput(), reset_locators=False) 732 self.pipeline = utils.OperationNode( 733 "extract_cells_on_plane", 734 parents=[self], 735 comment=f"#cells {self.dataset.GetNumberOfCells()}", 736 c="#9e2a2b", 737 ) 738 return self 739 740 def extract_cells_on_sphere(self, center: tuple, radius: tuple) -> Self: 741 """ 742 Extract cells that are lying of the specified surface. 743 """ 744 bf = vtki.new("3DLinearGridCrinkleExtractor") 745 bf.SetInputData(self.dataset) 746 bf.CopyPointDataOn() 747 bf.CopyCellDataOn() 748 bf.RemoveUnusedPointsOff() 749 750 sph = vtki.new("Sphere") 751 sph.SetRadius(radius) 752 sph.SetCenter(center) 753 bf.SetImplicitFunction(sph) 754 bf.Update() 755 756 self._update(bf.GetOutput()) 757 self.pipeline = utils.OperationNode( 758 "extract_cells_on_sphere", 759 parents=[self], 760 comment=f"#cells {self.dataset.GetNumberOfCells()}", 761 c="#9e2a2b", 762 ) 763 return self 764 765 def extract_cells_on_cylinder(self, center: tuple, axis: tuple, radius: float) -> Self: 766 """ 767 Extract cells that are lying of the specified surface. 768 """ 769 bf = vtki.new("3DLinearGridCrinkleExtractor") 770 bf.SetInputData(self.dataset) 771 bf.CopyPointDataOn() 772 bf.CopyCellDataOn() 773 bf.RemoveUnusedPointsOff() 774 775 cyl = vtki.new("Cylinder") 776 cyl.SetRadius(radius) 777 cyl.SetCenter(center) 778 cyl.SetAxis(axis) 779 bf.SetImplicitFunction(cyl) 780 bf.Update() 781 782 self.pipeline = utils.OperationNode( 783 "extract_cells_on_cylinder", 784 parents=[self], 785 comment=f"#cells {self.dataset.GetNumberOfCells()}", 786 c="#9e2a2b", 787 ) 788 self._update(bf.GetOutput()) 789 return self 790 791 def cut_with_plane(self, origin=(0, 0, 0), normal="x") -> "UnstructuredGrid": 792 """ 793 Cut the object with the plane defined by a point and a normal. 794 795 Arguments: 796 origin : (list) 797 the cutting plane goes through this point 798 normal : (list, str) 799 normal vector to the cutting plane 800 """ 801 # if isinstance(self, vedo.Volume): 802 # raise RuntimeError("cut_with_plane() is not applicable to Volume objects.") 803 804 strn = str(normal) 805 if strn == "x": normal = (1, 0, 0) 806 elif strn == "y": normal = (0, 1, 0) 807 elif strn == "z": normal = (0, 0, 1) 808 elif strn == "-x": normal = (-1, 0, 0) 809 elif strn == "-y": normal = (0, -1, 0) 810 elif strn == "-z": normal = (0, 0, -1) 811 plane = vtki.new("Plane") 812 plane.SetOrigin(origin) 813 plane.SetNormal(normal) 814 clipper = vtki.new("ClipDataSet") 815 clipper.SetInputData(self.dataset) 816 clipper.SetClipFunction(plane) 817 clipper.GenerateClipScalarsOff() 818 clipper.GenerateClippedOutputOff() 819 clipper.SetValue(0) 820 clipper.Update() 821 cout = clipper.GetOutput() 822 823 if isinstance(cout, vtki.vtkUnstructuredGrid): 824 ug = vedo.UnstructuredGrid(cout) 825 if isinstance(self, vedo.UnstructuredGrid): 826 self._update(cout) 827 self.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 828 return self 829 ug.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 830 return ug 831 832 else: 833 self._update(cout) 834 self.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 835 return self 836 837 def cut_with_box(self, box: Any) -> "UnstructuredGrid": 838 """ 839 Cut the grid with the specified bounding box. 840 841 Parameter box has format [xmin, xmax, ymin, ymax, zmin, zmax]. 842 If an object is passed, its bounding box are used. 843 844 This method always returns a TetMesh object. 845 846 Example: 847 ```python 848 from vedo import * 849 tmesh = TetMesh(dataurl+'limb_ugrid.vtk') 850 tmesh.color('rainbow') 851 cu = Cube(side=500).x(500) # any Mesh works 852 tmesh.cut_with_box(cu).show(axes=1) 853 ``` 854 855 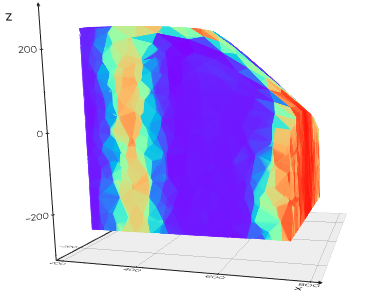 856 """ 857 bc = vtki.new("BoxClipDataSet") 858 bc.SetInputData(self.dataset) 859 try: 860 boxb = box.bounds() 861 except AttributeError: 862 boxb = box 863 864 bc.SetBoxClip(*boxb) 865 bc.Update() 866 cout = bc.GetOutput() 867 868 # output of vtkBoxClipDataSet is always tetrahedrons 869 tm = vedo.TetMesh(cout) 870 tm.pipeline = utils.OperationNode("cut_with_box", parents=[self], c="#9e2a2b") 871 return tm 872 873 def cut_with_mesh(self, mesh: Mesh, invert=False, whole_cells=False, on_boundary=False) -> "UnstructuredGrid": 874 """ 875 Cut a `UnstructuredGrid` or `TetMesh` with a `Mesh`. 876 877 Use `invert` to return cut off part of the input object. 878 """ 879 ug = self.dataset 880 881 ippd = vtki.new("ImplicitPolyDataDistance") 882 ippd.SetInput(mesh.dataset) 883 884 if whole_cells or on_boundary: 885 clipper = vtki.new("ExtractGeometry") 886 clipper.SetInputData(ug) 887 clipper.SetImplicitFunction(ippd) 888 clipper.SetExtractInside(not invert) 889 clipper.SetExtractBoundaryCells(False) 890 if on_boundary: 891 clipper.SetExtractBoundaryCells(True) 892 clipper.SetExtractOnlyBoundaryCells(True) 893 else: 894 signed_dists = vtki.vtkFloatArray() 895 signed_dists.SetNumberOfComponents(1) 896 signed_dists.SetName("SignedDistance") 897 for pointId in range(ug.GetNumberOfPoints()): 898 p = ug.GetPoint(pointId) 899 signed_dist = ippd.EvaluateFunction(p) 900 signed_dists.InsertNextValue(signed_dist) 901 ug.GetPointData().AddArray(signed_dists) 902 ug.GetPointData().SetActiveScalars("SignedDistance") # NEEDED 903 clipper = vtki.new("ClipDataSet") 904 clipper.SetInputData(ug) 905 clipper.SetInsideOut(not invert) 906 clipper.SetValue(0.0) 907 908 clipper.Update() 909 910 out = vedo.UnstructuredGrid(clipper.GetOutput()) 911 out.pipeline = utils.OperationNode("cut_with_mesh", parents=[self], c="#9e2a2b") 912 return out 913 914 915########################################################################## 916class TetMesh(UnstructuredGrid): 917 """The class describing tetrahedral meshes.""" 918 919 def __init__(self, inputobj=None): 920 """ 921 Arguments: 922 inputobj : (vtkUnstructuredGrid, list, str, tetgenpy.TetgenIO) 923 list of points and tet indices, or filename 924 """ 925 super().__init__() 926 927 self.dataset = None 928 929 self.mapper = vtki.new("PolyDataMapper") 930 self._actor = vtki.vtkActor() 931 self._actor.retrieve_object = weak_ref_to(self) 932 self._actor.SetMapper(self.mapper) 933 self.properties = self._actor.GetProperty() 934 935 self.name = "TetMesh" 936 937 # print('TetMesh inputtype', type(inputobj)) 938 939 ################### 940 if inputobj is None: 941 self.dataset = vtki.vtkUnstructuredGrid() 942 943 elif isinstance(inputobj, vtki.vtkUnstructuredGrid): 944 self.dataset = inputobj 945 946 elif isinstance(inputobj, UnstructuredGrid): 947 self.dataset = inputobj.dataset 948 949 elif "TetgenIO" in str(type(inputobj)): # tetgenpy object 950 inputobj = [inputobj.points(), inputobj.tetrahedra()] 951 952 elif isinstance(inputobj, vtki.vtkRectilinearGrid): 953 r2t = vtki.new("RectilinearGridToTetrahedra") 954 r2t.SetInputData(inputobj) 955 r2t.RememberVoxelIdOn() 956 r2t.SetTetraPerCellTo6() 957 r2t.Update() 958 self.dataset = r2t.GetOutput() 959 960 elif isinstance(inputobj, vtki.vtkDataSet): 961 r2t = vtki.new("DataSetTriangleFilter") 962 r2t.SetInputData(inputobj) 963 r2t.TetrahedraOnlyOn() 964 r2t.Update() 965 self.dataset = r2t.GetOutput() 966 967 elif isinstance(inputobj, str): 968 if "https://" in inputobj: 969 inputobj = download(inputobj, verbose=False) 970 if inputobj.endswith(".vtu"): 971 reader = vtki.new("XMLUnstructuredGridReader") 972 else: 973 reader = vtki.new("UnstructuredGridReader") 974 975 if not os.path.isfile(inputobj): 976 # for some reason vtk Reader does not complain 977 vedo.logger.error(f"file {inputobj} not found") 978 raise FileNotFoundError 979 980 self.filename = inputobj 981 reader.SetFileName(inputobj) 982 reader.Update() 983 ug = reader.GetOutput() 984 985 tt = vtki.new("DataSetTriangleFilter") 986 tt.SetInputData(ug) 987 tt.SetTetrahedraOnly(True) 988 tt.Update() 989 self.dataset = tt.GetOutput() 990 991 ############################### 992 if utils.is_sequence(inputobj): 993 self.dataset = vtki.vtkUnstructuredGrid() 994 995 points, cells = inputobj 996 if len(points) == 0: 997 return 998 if not utils.is_sequence(points[0]): 999 return 1000 if len(cells) == 0: 1001 return 1002 1003 if not utils.is_sequence(cells[0]): 1004 tets = [] 1005 nf = cells[0] + 1 1006 for i, cl in enumerate(cells): 1007 if i in (nf, 0): 1008 k = i + 1 1009 nf = cl + k 1010 cell = [cells[j + k] for j in range(cl)] 1011 tets.append(cell) 1012 cells = tets 1013 1014 source_points = vtki.vtkPoints() 1015 varr = utils.numpy2vtk(points, dtype=np.float32) 1016 source_points.SetData(varr) 1017 self.dataset.SetPoints(source_points) 1018 1019 source_tets = vtki.vtkCellArray() 1020 for f in cells: 1021 ele = vtki.vtkTetra() 1022 pid = ele.GetPointIds() 1023 for i, fi in enumerate(f): 1024 pid.SetId(i, fi) 1025 source_tets.InsertNextCell(ele) 1026 self.dataset.SetCells(vtki.cell_types["TETRA"], source_tets) 1027 1028 if not self.dataset: 1029 vedo.logger.error(f"cannot understand input type {type(inputobj)}") 1030 return 1031 1032 self.properties.SetColor(0.352, 0.612, 0.996) # blue7 1033 1034 self.pipeline = utils.OperationNode( 1035 self, comment=f"#tets {self.dataset.GetNumberOfCells()}", c="#9e2a2b" 1036 ) 1037 1038 ################################################################## 1039 def __str__(self): 1040 """Print a string summary of the `TetMesh` object.""" 1041 module = self.__class__.__module__ 1042 name = self.__class__.__name__ 1043 out = vedo.printc( 1044 f"{module}.{name} at ({hex(self.memory_address())})".ljust(75), 1045 c="c", bold=True, invert=True, return_string=True, 1046 ) 1047 out += "\x1b[0m\u001b[36m" 1048 1049 out += "nr. of verts".ljust(14) + ": " + str(self.npoints) + "\n" 1050 out += "nr. of tetras".ljust(14) + ": " + str(self.ncells) + "\n" 1051 1052 if self.npoints: 1053 out+="size".ljust(14)+ ": average=" + utils.precision(self.average_size(),6) 1054 out+=", diagonal="+ utils.precision(self.diagonal_size(), 6)+ "\n" 1055 out+="center of mass".ljust(14) + ": " + utils.precision(self.center_of_mass(),6)+"\n" 1056 1057 bnds = self.bounds() 1058 bx1, bx2 = utils.precision(bnds[0], 3), utils.precision(bnds[1], 3) 1059 by1, by2 = utils.precision(bnds[2], 3), utils.precision(bnds[3], 3) 1060 bz1, bz2 = utils.precision(bnds[4], 3), utils.precision(bnds[5], 3) 1061 out += "bounds".ljust(14) + ":" 1062 out += " x=(" + bx1 + ", " + bx2 + ")," 1063 out += " y=(" + by1 + ", " + by2 + ")," 1064 out += " z=(" + bz1 + ", " + bz2 + ")\n" 1065 1066 for key in self.pointdata.keys(): 1067 arr = self.pointdata[key] 1068 rng = utils.precision(arr.min(), 3) + ", " + utils.precision(arr.max(), 3) 1069 mark_active = "pointdata" 1070 a_scalars = self.dataset.GetPointData().GetScalars() 1071 a_vectors = self.dataset.GetPointData().GetVectors() 1072 a_tensors = self.dataset.GetPointData().GetTensors() 1073 if a_scalars and a_scalars.GetName() == key: 1074 mark_active += " *" 1075 elif a_vectors and a_vectors.GetName() == key: 1076 mark_active += " **" 1077 elif a_tensors and a_tensors.GetName() == key: 1078 mark_active += " ***" 1079 out += mark_active.ljust(14) + f': "{key}" ({arr.dtype}), ndim={arr.ndim}' 1080 out += f", range=({rng})\n" 1081 1082 for key in self.celldata.keys(): 1083 arr = self.celldata[key] 1084 rng = utils.precision(arr.min(), 3) + ", " + utils.precision(arr.max(), 3) 1085 mark_active = "celldata" 1086 a_scalars = self.dataset.GetCellData().GetScalars() 1087 a_vectors = self.dataset.GetCellData().GetVectors() 1088 a_tensors = self.dataset.GetCellData().GetTensors() 1089 if a_scalars and a_scalars.GetName() == key: 1090 mark_active += " *" 1091 elif a_vectors and a_vectors.GetName() == key: 1092 mark_active += " **" 1093 elif a_tensors and a_tensors.GetName() == key: 1094 mark_active += " ***" 1095 out += mark_active.ljust(14) + f': "{key}" ({arr.dtype}), ndim={arr.ndim}' 1096 out += f", range=({rng})\n" 1097 1098 for key in self.metadata.keys(): 1099 arr = self.metadata[key] 1100 out += "metadata".ljust(14) + ": " + f'"{key}" ({len(arr)} values)\n' 1101 1102 return out.rstrip() + "\x1b[0m" 1103 1104 def _repr_html_(self): 1105 """ 1106 HTML representation of the TetMesh object for Jupyter Notebooks. 1107 1108 Returns: 1109 HTML text with the image and some properties. 1110 """ 1111 import io 1112 import base64 1113 from PIL import Image 1114 1115 library_name = "vedo.grids.TetMesh" 1116 help_url = "https://vedo.embl.es/docs/vedo/grids.html#TetMesh" 1117 1118 arr = self.thumbnail() 1119 im = Image.fromarray(arr) 1120 buffered = io.BytesIO() 1121 im.save(buffered, format="PNG", quality=100) 1122 encoded = base64.b64encode(buffered.getvalue()).decode("utf-8") 1123 url = "data:image/png;base64," + encoded 1124 image = f"<img src='{url}'></img>" 1125 1126 bounds = "<br/>".join( 1127 [ 1128 utils.precision(min_x, 4) + " ... " + utils.precision(max_x, 4) 1129 for min_x, max_x in zip(self.bounds()[::2], self.bounds()[1::2]) 1130 ] 1131 ) 1132 1133 help_text = "" 1134 if self.name: 1135 help_text += f"<b> {self.name}:   </b>" 1136 help_text += '<b><a href="' + help_url + '" target="_blank">' + library_name + "</a></b>" 1137 if self.filename: 1138 dots = "" 1139 if len(self.filename) > 30: 1140 dots = "..." 1141 help_text += f"<br/><code><i>({dots}{self.filename[-30:]})</i></code>" 1142 1143 pdata = "" 1144 if self.dataset.GetPointData().GetScalars(): 1145 if self.dataset.GetPointData().GetScalars().GetName(): 1146 name = self.dataset.GetPointData().GetScalars().GetName() 1147 pdata = "<tr><td><b> point data array </b></td><td>" + name + "</td></tr>" 1148 1149 cdata = "" 1150 if self.dataset.GetCellData().GetScalars(): 1151 if self.dataset.GetCellData().GetScalars().GetName(): 1152 name = self.dataset.GetCellData().GetScalars().GetName() 1153 cdata = "<tr><td><b> cell data array </b></td><td>" + name + "</td></tr>" 1154 1155 pts = self.coordinates 1156 cm = np.mean(pts, axis=0) 1157 1158 allt = [ 1159 "<table>", 1160 "<tr>", 1161 "<td>", image, "</td>", 1162 "<td style='text-align: center; vertical-align: center;'><br/>", help_text, 1163 "<table>", 1164 "<tr><td><b> bounds </b> <br/> (x/y/z) </td><td>" + str(bounds) + "</td></tr>", 1165 "<tr><td><b> center of mass </b></td><td>" + utils.precision(cm,3) + "</td></tr>", 1166 "<tr><td><b> nr. points / tets </b></td><td>" 1167 + str(self.npoints) + " / " + str(self.ncells) + "</td></tr>", 1168 pdata, 1169 cdata, 1170 "</table>", 1171 "</table>", 1172 ] 1173 return "\n".join(allt) 1174 1175 def compute_quality(self, metric=7) -> np.ndarray: 1176 """ 1177 Calculate functions of quality for the elements of a tetrahedral mesh. 1178 This method adds to the mesh a cell array named "Quality". 1179 1180 Arguments: 1181 metric : (int) 1182 type of estimators: 1183 - EDGE RATIO, 0 1184 - ASPECT RATIO, 1 1185 - RADIUS RATIO, 2 1186 - ASPECT FROBENIUS, 3 1187 - MIN_ANGLE, 4 1188 - COLLAPSE RATIO, 5 1189 - ASPECT GAMMA, 6 1190 - VOLUME, 7 1191 1192 See class [vtkMeshQuality](https://vtk.org/doc/nightly/html/classvtkMeshQuality.html) 1193 for an explanation of the meaning of each metric.. 1194 """ 1195 qf = vtki.new("MeshQuality") 1196 qf.SetInputData(self.dataset) 1197 qf.SetTetQualityMeasure(metric) 1198 qf.SaveCellQualityOn() 1199 qf.Update() 1200 self._update(qf.GetOutput()) 1201 return utils.vtk2numpy(qf.GetOutput().GetCellData().GetArray("Quality")) 1202 1203 def check_validity(self, tol=0) -> np.ndarray: 1204 """ 1205 Return an array of possible problematic tets following this convention: 1206 ```python 1207 Valid = 0 1208 WrongNumberOfPoints = 01 1209 IntersectingEdges = 02 1210 IntersectingFaces = 04 1211 NoncontiguousEdges = 08 1212 Nonconvex = 10 1213 OrientedIncorrectly = 20 1214 ``` 1215 1216 Arguments: 1217 tol : (float) 1218 This value is used as an epsilon for floating point 1219 equality checks throughout the cell checking process. 1220 """ 1221 vald = vtki.new("CellValidator") 1222 if tol: 1223 vald.SetTolerance(tol) 1224 vald.SetInputData(self.dataset) 1225 vald.Update() 1226 varr = vald.GetOutput().GetCellData().GetArray("ValidityState") 1227 return utils.vtk2numpy(varr) 1228 1229 def decimate(self, scalars_name: str, fraction=0.5, n=0) -> Self: 1230 """ 1231 Downsample the number of tets in a TetMesh to a specified fraction. 1232 Either `fraction` or `n` must be set. 1233 1234 Arguments: 1235 fraction : (float) 1236 the desired final fraction of the total. 1237 n : (int) 1238 the desired number of final tets 1239 1240 .. note:: setting `fraction=0.1` leaves 10% of the original nr of tets. 1241 """ 1242 decimate = vtki.new("UnstructuredGridQuadricDecimation") 1243 decimate.SetInputData(self.dataset) 1244 decimate.SetScalarsName(scalars_name) 1245 1246 if n: # n = desired number of points 1247 decimate.SetNumberOfTetsOutput(n) 1248 else: 1249 decimate.SetTargetReduction(1 - fraction) 1250 decimate.Update() 1251 self._update(decimate.GetOutput()) 1252 self.pipeline = utils.OperationNode( 1253 "decimate", comment=f"array: {scalars_name}", c="#edabab", parents=[self] 1254 ) 1255 return self 1256 1257 def subdivide(self) -> Self: 1258 """ 1259 Increase the number of tetrahedrons of a `TetMesh`. 1260 Subdivides each tetrahedron into twelve smaller tetras. 1261 """ 1262 sd = vtki.new("SubdivideTetra") 1263 sd.SetInputData(self.dataset) 1264 sd.Update() 1265 self._update(sd.GetOutput()) 1266 self.pipeline = utils.OperationNode("subdivide", c="#edabab", parents=[self]) 1267 return self 1268 1269 def generate_random_points(self, n, min_radius=0) -> "vedo.Points": 1270 """ 1271 Generate `n` uniformly distributed random points 1272 inside the tetrahedral mesh. 1273 1274 A new point data array is added to the output points 1275 called "OriginalCellID" which contains the index of 1276 the cell ID in which the point was generated. 1277 1278 Arguments: 1279 n : (int) 1280 number of points to generate. 1281 min_radius: (float) 1282 impose a minimum distance between points. 1283 If `min_radius` is set to 0, the points are 1284 generated uniformly at random inside the mesh. 1285 If `min_radius` is set to a positive value, 1286 the points are generated uniformly at random 1287 inside the mesh, but points closer than `min_radius` 1288 to any other point are discarded. 1289 1290 Returns a `vedo.Points` object. 1291 1292 Note: 1293 Consider using `points.probe(msh)` to interpolate 1294 any existing mesh data onto the points. 1295 1296 Example: 1297 ```python 1298 from vedo import * 1299 tmesh = TetMesh(dataurl + "limb.vtu").alpha(0.2) 1300 pts = tmesh.generate_random_points(20000, min_radius=10) 1301 print(pts.pointdata["OriginalCellID"]) 1302 show(pts, tmesh, axes=1).close() 1303 ``` 1304 """ 1305 cmesh = self.compute_cell_size() 1306 tets = cmesh.cells 1307 verts = cmesh.coordinates 1308 cumul = np.cumsum(np.abs(cmesh.celldata["Volume"])) 1309 1310 out_pts = [] 1311 orig_cell = [] 1312 for _ in range(n): 1313 random_area = np.random.random() * cumul[-1] 1314 it = np.searchsorted(cumul, random_area) 1315 A, B, C, D = verts[tets[it]] 1316 r1, r2, r3 = sorted(np.random.random(3)) 1317 p = r1 * A + (r2 - r1) * B + (r3 - r2) * C + (1 - r3) * D 1318 out_pts.append(p) 1319 orig_cell.append(it) 1320 orig_cellnp = np.array(orig_cell, dtype=np.uint32) 1321 1322 vpts = vedo.pointcloud.Points(out_pts) 1323 vpts.pointdata["OriginalCellID"] = orig_cellnp 1324 1325 if min_radius > 0: 1326 vpts.subsample(min_radius, absolute=True) 1327 1328 vpts.point_size(5).color("k1") 1329 vpts.name = "RandomPoints" 1330 vpts.pipeline = utils.OperationNode( 1331 "generate_random_points", c="#edabab", parents=[self]) 1332 return vpts 1333 1334 def isosurface(self, value=True, flying_edges=None) -> "vedo.Mesh": 1335 """ 1336 Return a `vedo.Mesh` isosurface. 1337 The "isosurface" is the surface of the region of points 1338 whose values equal to `value`. 1339 1340 Set `value` to a single value or list of values to compute the isosurface(s). 1341 1342 Note that flying_edges option is not available for `TetMesh`. 1343 """ 1344 if flying_edges is not None: 1345 vedo.logger.warning("flying_edges option is not available for TetMesh.") 1346 1347 if not self.dataset.GetPointData().GetScalars(): 1348 vedo.logger.warning( 1349 "in isosurface() no scalar pointdata found. " 1350 "Mappping cells to points." 1351 ) 1352 self.map_cells_to_points() 1353 scrange = self.dataset.GetPointData().GetScalars().GetRange() 1354 cf = vtki.new("ContourFilter") # vtki.new("ContourGrid") 1355 cf.SetInputData(self.dataset) 1356 1357 if utils.is_sequence(value): 1358 cf.SetNumberOfContours(len(value)) 1359 for i, t in enumerate(value): 1360 cf.SetValue(i, t) 1361 cf.Update() 1362 else: 1363 if value is True: 1364 value = (2 * scrange[0] + scrange[1]) / 3.0 1365 cf.SetValue(0, value) 1366 cf.Update() 1367 1368 msh = Mesh(cf.GetOutput(), c=None) 1369 msh.copy_properties_from(self) 1370 msh.pipeline = utils.OperationNode("isosurface", c="#edabab", parents=[self]) 1371 return msh 1372 1373 def slice(self, origin=(0, 0, 0), normal=(1, 0, 0)) -> "vedo.Mesh": 1374 """ 1375 Return a 2D slice of the mesh by a plane passing through origin and assigned normal. 1376 """ 1377 strn = str(normal) 1378 if strn == "x": normal = (1, 0, 0) 1379 elif strn == "y": normal = (0, 1, 0) 1380 elif strn == "z": normal = (0, 0, 1) 1381 elif strn == "-x": normal = (-1, 0, 0) 1382 elif strn == "-y": normal = (0, -1, 0) 1383 elif strn == "-z": normal = (0, 0, -1) 1384 plane = vtki.new("Plane") 1385 plane.SetOrigin(origin) 1386 plane.SetNormal(normal) 1387 1388 cc = vtki.new("Cutter") 1389 cc.SetInputData(self.dataset) 1390 cc.SetCutFunction(plane) 1391 cc.Update() 1392 msh = Mesh(cc.GetOutput()).flat().lighting("ambient") 1393 msh.copy_properties_from(self) 1394 msh.pipeline = utils.OperationNode("slice", c="#edabab", parents=[self]) 1395 return msh 1396 1397 1398########################################################################## 1399class RectilinearGrid(PointAlgorithms, MeshVisual): 1400 """ 1401 Build a rectilinear grid. 1402 """ 1403 1404 def __init__(self, inputobj=None): 1405 """ 1406 A RectilinearGrid is a dataset where edges are parallel to the coordinate axes. 1407 It can be thought of as a tessellation of a box in 3D space, similar to a `Volume` 1408 except that the cells are not necessarily cubes, but they can have different lengths 1409 along each axis. 1410 This can be useful to describe a volume with variable resolution where one needs 1411 to represent a region with higher detail with respect to another region. 1412 1413 Arguments: 1414 inputobj : (vtkRectilinearGrid, list, str) 1415 list of points and indices, or filename 1416 1417 Example: 1418 ```python 1419 from vedo import RectilinearGrid, show 1420 1421 xcoords = 7 + np.sqrt(np.arange(0,2500,25)) 1422 ycoords = np.arange(0, 20) 1423 zcoords = np.arange(0, 20) 1424 1425 rgrid = RectilinearGrid([xcoords, ycoords, zcoords]) 1426 1427 print(rgrid) 1428 print(rgrid.x_coordinates().shape) 1429 print(rgrid.compute_structured_coords([20,10,11])) 1430 1431 msh = rgrid.tomesh().lw(1) 1432 1433 show(msh, axes=1, viewup="z") 1434 ``` 1435 """ 1436 1437 super().__init__() 1438 1439 self.dataset = None 1440 1441 self.mapper = vtki.new("PolyDataMapper") 1442 self._actor = vtki.vtkActor() 1443 self._actor.retrieve_object = weak_ref_to(self) 1444 self._actor.SetMapper(self.mapper) 1445 self.properties = self._actor.GetProperty() 1446 1447 self.transform = LinearTransform() 1448 self.point_locator = None 1449 self.cell_locator = None 1450 self.line_locator = None 1451 1452 self.name = "RectilinearGrid" 1453 self.filename = "" 1454 1455 self.info = {} 1456 self.time = time.time() 1457 1458 ############################### 1459 if inputobj is None: 1460 self.dataset = vtki.vtkRectilinearGrid() 1461 1462 elif isinstance(inputobj, vtki.vtkRectilinearGrid): 1463 self.dataset = inputobj 1464 1465 elif isinstance(inputobj, RectilinearGrid): 1466 self.dataset = inputobj.dataset 1467 1468 elif isinstance(inputobj, str): 1469 if "https://" in inputobj: 1470 inputobj = download(inputobj, verbose=False) 1471 if inputobj.endswith(".vtr"): 1472 reader = vtki.new("XMLRectilinearGridReader") 1473 else: 1474 reader = vtki.new("RectilinearGridReader") 1475 self.filename = inputobj 1476 reader.SetFileName(inputobj) 1477 reader.Update() 1478 self.dataset = reader.GetOutput() 1479 1480 elif utils.is_sequence(inputobj): 1481 self.dataset = vtki.vtkRectilinearGrid() 1482 xcoords, ycoords, zcoords = inputobj 1483 nx, ny, nz = len(xcoords), len(ycoords), len(zcoords) 1484 self.dataset.SetDimensions(nx, ny, nz) 1485 self.dataset.SetXCoordinates(utils.numpy2vtk(xcoords)) 1486 self.dataset.SetYCoordinates(utils.numpy2vtk(ycoords)) 1487 self.dataset.SetZCoordinates(utils.numpy2vtk(zcoords)) 1488 1489 ############################### 1490 1491 if not self.dataset: 1492 vedo.logger.error(f"RectilinearGrid: cannot understand input type {type(inputobj)}") 1493 return 1494 1495 self.properties.SetColor(0.352, 0.612, 0.996) # blue7 1496 1497 self.pipeline = utils.OperationNode( 1498 self, comment=f"#tets {self.dataset.GetNumberOfCells()}", c="#9e2a2b" 1499 ) 1500 1501 @property 1502 def actor(self): 1503 """Return the `vtkActor` of the object.""" 1504 gf = vtki.new("GeometryFilter") 1505 gf.SetInputData(self.dataset) 1506 gf.Update() 1507 self.mapper.SetInputData(gf.GetOutput()) 1508 self.mapper.Modified() 1509 return self._actor 1510 1511 @actor.setter 1512 def actor(self, _): 1513 pass 1514 1515 def _update(self, data, reset_locators=False): 1516 self.dataset = data 1517 if reset_locators: 1518 self.cell_locator = None 1519 self.point_locator = None 1520 return self 1521 1522 ################################################################## 1523 def __str__(self): 1524 """Print a summary for the `RectilinearGrid` object.""" 1525 module = self.__class__.__module__ 1526 name = self.__class__.__name__ 1527 out = vedo.printc( 1528 f"{module}.{name} at ({hex(self.memory_address())})".ljust(75), 1529 c="c", bold=True, invert=True, return_string=True, 1530 ) 1531 out += "\x1b[0m\x1b[36;1m" 1532 1533 out += "name".ljust(14) + ": " + str(self.name) + "\n" 1534 if self.filename: 1535 out += "filename".ljust(14) + ": " + str(self.filename) + "\n" 1536 1537 out += "dimensions".ljust(14) + ": " + str(self.dataset.GetDimensions()) + "\n" 1538 1539 out += "center".ljust(14) + ": " 1540 out += utils.precision(self.dataset.GetCenter(), 6) + "\n" 1541 1542 bnds = self.bounds() 1543 bx1, bx2 = utils.precision(bnds[0], 3), utils.precision(bnds[1], 3) 1544 by1, by2 = utils.precision(bnds[2], 3), utils.precision(bnds[3], 3) 1545 bz1, bz2 = utils.precision(bnds[4], 3), utils.precision(bnds[5], 3) 1546 out += "bounds".ljust(14) + ":" 1547 out += " x=(" + bx1 + ", " + bx2 + ")," 1548 out += " y=(" + by1 + ", " + by2 + ")," 1549 out += " z=(" + bz1 + ", " + bz2 + ")\n" 1550 1551 out += "memory size".ljust(14) + ": " 1552 out += utils.precision(self.dataset.GetActualMemorySize() / 1024, 2) + " MB\n" 1553 1554 for key in self.pointdata.keys(): 1555 arr = self.pointdata[key] 1556 rng = utils.precision(arr.min(), 3) + ", " + utils.precision(arr.max(), 3) 1557 mark_active = "pointdata" 1558 a_scalars = self.dataset.GetPointData().GetScalars() 1559 a_vectors = self.dataset.GetPointData().GetVectors() 1560 a_tensors = self.dataset.GetPointData().GetTensors() 1561 if a_scalars and a_scalars.GetName() == key: 1562 mark_active += " *" 1563 elif a_vectors and a_vectors.GetName() == key: 1564 mark_active += " **" 1565 elif a_tensors and a_tensors.GetName() == key: 1566 mark_active += " ***" 1567 out += mark_active.ljust(14) + f': "{key}" ({arr.dtype}), ndim={arr.ndim}' 1568 out += f", range=({rng})\n" 1569 1570 for key in self.celldata.keys(): 1571 arr = self.celldata[key] 1572 rng = utils.precision(arr.min(), 3) + ", " + utils.precision(arr.max(), 3) 1573 mark_active = "celldata" 1574 a_scalars = self.dataset.GetCellData().GetScalars() 1575 a_vectors = self.dataset.GetCellData().GetVectors() 1576 a_tensors = self.dataset.GetCellData().GetTensors() 1577 if a_scalars and a_scalars.GetName() == key: 1578 mark_active += " *" 1579 elif a_vectors and a_vectors.GetName() == key: 1580 mark_active += " **" 1581 elif a_tensors and a_tensors.GetName() == key: 1582 mark_active += " ***" 1583 out += mark_active.ljust(14) + f': "{key}" ({arr.dtype}), ndim={arr.ndim}' 1584 out += f", range=({rng})\n" 1585 1586 for key in self.metadata.keys(): 1587 arr = self.metadata[key] 1588 out += "metadata".ljust(14) + ": " + f'"{key}" ({len(arr)} values)\n' 1589 1590 return out.rstrip() + "\x1b[0m" 1591 1592 def _repr_html_(self): 1593 """ 1594 HTML representation of the RectilinearGrid object for Jupyter Notebooks. 1595 1596 Returns: 1597 HTML text with the image and some properties. 1598 """ 1599 import io 1600 import base64 1601 from PIL import Image 1602 1603 library_name = "vedo.grids.RectilinearGrid" 1604 help_url = "https://vedo.embl.es/docs/vedo/grids.html#RectilinearGrid" 1605 1606 m = self.tomesh().linewidth(1).lighting("off") 1607 arr= m.thumbnail(zoom=1, elevation=-30, azimuth=-30) 1608 1609 im = Image.fromarray(arr) 1610 buffered = io.BytesIO() 1611 im.save(buffered, format="PNG", quality=100) 1612 encoded = base64.b64encode(buffered.getvalue()).decode("utf-8") 1613 url = "data:image/png;base64," + encoded 1614 image = f"<img src='{url}'></img>" 1615 1616 bounds = "<br/>".join( 1617 [ 1618 utils.precision(min_x, 4) + " ... " + utils.precision(max_x, 4) 1619 for min_x, max_x in zip(self.bounds()[::2], self.bounds()[1::2]) 1620 ] 1621 ) 1622 1623 help_text = "" 1624 if self.name: 1625 help_text += f"<b> {self.name}:   </b>" 1626 help_text += '<b><a href="' + help_url + '" target="_blank">' + library_name + "</a></b>" 1627 if self.filename: 1628 dots = "" 1629 if len(self.filename) > 30: 1630 dots = "..." 1631 help_text += f"<br/><code><i>({dots}{self.filename[-30:]})</i></code>" 1632 1633 pdata = "" 1634 if self.dataset.GetPointData().GetScalars(): 1635 if self.dataset.GetPointData().GetScalars().GetName(): 1636 name = self.dataset.GetPointData().GetScalars().GetName() 1637 pdata = "<tr><td><b> point data array </b></td><td>" + name + "</td></tr>" 1638 1639 cdata = "" 1640 if self.dataset.GetCellData().GetScalars(): 1641 if self.dataset.GetCellData().GetScalars().GetName(): 1642 name = self.dataset.GetCellData().GetScalars().GetName() 1643 cdata = "<tr><td><b> cell data array </b></td><td>" + name + "</td></tr>" 1644 1645 pts = self.coordinates 1646 cm = np.mean(pts, axis=0) 1647 1648 _all = [ 1649 "<table>", 1650 "<tr>", 1651 "<td>", image, "</td>", 1652 "<td style='text-align: center; vertical-align: center;'><br/>", help_text, 1653 "<table>", 1654 "<tr><td><b> bounds </b> <br/> (x/y/z) </td><td>" + str(bounds) + "</td></tr>", 1655 "<tr><td><b> center of mass </b></td><td>" + utils.precision(cm,3) + "</td></tr>", 1656 "<tr><td><b> nr. points / cells </b></td><td>" 1657 + str(self.npoints) + " / " + str(self.ncells) + "</td></tr>", 1658 pdata, 1659 cdata, 1660 "</table>", 1661 "</table>", 1662 ] 1663 return "\n".join(_all) 1664 1665 def dimensions(self) -> np.ndarray: 1666 """Return the number of points in the x, y and z directions.""" 1667 return np.array(self.dataset.GetDimensions()) 1668 1669 def x_coordinates(self) -> np.ndarray: 1670 """Return the x-coordinates of the grid.""" 1671 return utils.vtk2numpy(self.dataset.GetXCoordinates()) 1672 1673 def y_coordinates(self) -> np.ndarray: 1674 """Return the y-coordinates of the grid.""" 1675 return utils.vtk2numpy(self.dataset.GetYCoordinates()) 1676 1677 def z_coordinates(self) -> np.ndarray: 1678 """Return the z-coordinates of the grid.""" 1679 return utils.vtk2numpy(self.dataset.GetZCoordinates()) 1680 1681 def is_point_visible(self, pid: int) -> bool: 1682 """Return True if point `pid` is visible.""" 1683 return self.dataset.IsPointVisible(pid) 1684 1685 def is_cell_visible(self, cid: int) -> bool: 1686 """Return True if cell `cid` is visible.""" 1687 return self.dataset.IsCellVisible(cid) 1688 1689 def has_blank_points(self) -> bool: 1690 """Return True if the grid has blank points.""" 1691 return self.dataset.HasAnyBlankPoints() 1692 1693 def has_blank_cells(self) -> bool: 1694 """Return True if the grid has blank cells.""" 1695 return self.dataset.HasAnyBlankCells() 1696 1697 def compute_structured_coords(self, x: list) -> dict: 1698 """ 1699 Convenience function computes the structured coordinates for a point `x`. 1700 1701 This method returns a dictionary with keys `ijk`, `pcoords` and `inside`. 1702 The cell is specified by the array `ijk`. 1703 and the parametric coordinates in the cell are specified with `pcoords`. 1704 Value of `inside` is False if the point x is outside of the grid. 1705 """ 1706 ijk = [0, 0, 0] 1707 pcoords = [0., 0., 0.] 1708 inout = self.dataset.ComputeStructuredCoordinates(x, ijk, pcoords) 1709 return {"ijk": np.array(ijk), "pcoords": np.array(pcoords), "inside": bool(inout)} 1710 1711 def compute_pointid(self, ijk: int) -> int: 1712 """Given a location in structured coordinates (i-j-k), return the point id.""" 1713 return self.dataset.ComputePointId(ijk) 1714 1715 def compute_cellid(self, ijk: int) -> int: 1716 """Given a location in structured coordinates (i-j-k), return the cell id.""" 1717 return self.dataset.ComputeCellId(ijk) 1718 1719 def find_point(self, x: list) -> int: 1720 """Given a position `x`, return the id of the closest point.""" 1721 return self.dataset.FindPoint(x) 1722 1723 def find_cell(self, x: list) -> dict: 1724 """Given a position `x`, return the id of the closest cell.""" 1725 cell = vtki.vtkHexagonalPrism() 1726 cellid = vtki.mutable(0) 1727 tol2 = 0.001 # vtki.mutable(0) 1728 subid = vtki.mutable(0) 1729 pcoords = [0.0, 0.0, 0.0] 1730 weights = [0.0, 0.0, 0.0] 1731 res = self.dataset.FindCell(x, cell, cellid, tol2, subid, pcoords, weights) 1732 result = {} 1733 result["cellid"] = cellid 1734 result["subid"] = subid 1735 result["pcoords"] = pcoords 1736 result["weights"] = weights 1737 result["status"] = res 1738 return result 1739 1740 def clone(self, deep=True) -> "RectilinearGrid": 1741 """Return a clone copy of the RectilinearGrid. Alias of `copy()`.""" 1742 if deep: 1743 newrg = vtki.vtkRectilinearGrid() 1744 newrg.CopyStructure(self.dataset) 1745 newrg.CopyAttributes(self.dataset) 1746 newvol = RectilinearGrid(newrg) 1747 else: 1748 newvol = RectilinearGrid(self.dataset) 1749 1750 prop = vtki.vtkProperty() 1751 prop.DeepCopy(self.properties) 1752 newvol.actor.SetProperty(prop) 1753 newvol.properties = prop 1754 newvol.pipeline = utils.OperationNode("clone", parents=[self], c="#bbd0ff", shape="diamond") 1755 return newvol 1756 1757 def bounds(self) -> np.ndarray: 1758 """ 1759 Get the object bounds. 1760 Returns a list in format `[xmin,xmax, ymin,ymax, zmin,zmax]`. 1761 """ 1762 # OVERRIDE CommonAlgorithms.bounds() which is too slow 1763 return np.array(self.dataset.GetBounds()) 1764 1765 def isosurface(self, value=None) -> "vedo.Mesh": 1766 """ 1767 Return a `Mesh` isosurface extracted from the object. 1768 1769 Set `value` as single float or list of values to draw the isosurface(s). 1770 """ 1771 scrange = self.dataset.GetScalarRange() 1772 1773 cf = vtki.new("ContourFilter") 1774 cf.UseScalarTreeOn() 1775 cf.SetInputData(self.dataset) 1776 cf.ComputeNormalsOn() 1777 1778 if value is None: 1779 value = (2 * scrange[0] + scrange[1]) / 3.0 1780 # print("automatic isosurface value =", value) 1781 cf.SetValue(0, value) 1782 else: 1783 if utils.is_sequence(value): 1784 cf.SetNumberOfContours(len(value)) 1785 for i, t in enumerate(value): 1786 cf.SetValue(i, t) 1787 else: 1788 cf.SetValue(0, value) 1789 1790 cf.Update() 1791 poly = cf.GetOutput() 1792 1793 out = vedo.mesh.Mesh(poly, c=None).phong() 1794 out.mapper.SetScalarRange(scrange[0], scrange[1]) 1795 1796 out.pipeline = utils.OperationNode( 1797 "isosurface", 1798 parents=[self], 1799 comment=f"#pts {out.dataset.GetNumberOfPoints()}", 1800 c="#4cc9f0:#e9c46a", 1801 ) 1802 return out 1803 1804 def cut_with_plane(self, origin=(0, 0, 0), normal="x") -> "UnstructuredGrid": 1805 """ 1806 Cut the object with the plane defined by a point and a normal. 1807 1808 Arguments: 1809 origin : (list) 1810 the cutting plane goes through this point 1811 normal : (list, str) 1812 normal vector to the cutting plane 1813 """ 1814 strn = str(normal) 1815 if strn == "x": normal = (1, 0, 0) 1816 elif strn == "y": normal = (0, 1, 0) 1817 elif strn == "z": normal = (0, 0, 1) 1818 elif strn == "-x": normal = (-1, 0, 0) 1819 elif strn == "-y": normal = (0, -1, 0) 1820 elif strn == "-z": normal = (0, 0, -1) 1821 plane = vtki.new("Plane") 1822 plane.SetOrigin(origin) 1823 plane.SetNormal(normal) 1824 clipper = vtki.new("ClipDataSet") 1825 clipper.SetInputData(self.dataset) 1826 clipper.SetClipFunction(plane) 1827 clipper.GenerateClipScalarsOff() 1828 clipper.GenerateClippedOutputOff() 1829 clipper.SetValue(0) 1830 clipper.Update() 1831 cout = clipper.GetOutput() 1832 ug = vedo.UnstructuredGrid(cout) 1833 if isinstance(self, UnstructuredGrid): 1834 self._update(cout) 1835 self.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 1836 return self 1837 ug.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 1838 return ug 1839 1840 def cut_with_mesh(self, mesh, invert=False, whole_cells=False, on_boundary=False) -> "UnstructuredGrid": 1841 """ 1842 Cut a `RectilinearGrid` with a `Mesh`. 1843 1844 Use `invert` to return cut off part of the input object. 1845 """ 1846 ug = self.dataset 1847 1848 ippd = vtki.new("ImplicitPolyDataDistance") 1849 ippd.SetInput(mesh.dataset) 1850 1851 if whole_cells or on_boundary: 1852 clipper = vtki.new("ExtractGeometry") 1853 clipper.SetInputData(ug) 1854 clipper.SetImplicitFunction(ippd) 1855 clipper.SetExtractInside(not invert) 1856 clipper.SetExtractBoundaryCells(False) 1857 if on_boundary: 1858 clipper.SetExtractBoundaryCells(True) 1859 clipper.SetExtractOnlyBoundaryCells(True) 1860 else: 1861 signed_dists = vtki.vtkFloatArray() 1862 signed_dists.SetNumberOfComponents(1) 1863 signed_dists.SetName("SignedDistance") 1864 for pointId in range(ug.GetNumberOfPoints()): 1865 p = ug.GetPoint(pointId) 1866 signed_dist = ippd.EvaluateFunction(p) 1867 signed_dists.InsertNextValue(signed_dist) 1868 ug.GetPointData().AddArray(signed_dists) 1869 ug.GetPointData().SetActiveScalars("SignedDistance") # NEEDED 1870 clipper = vtki.new("ClipDataSet") 1871 clipper.SetInputData(ug) 1872 clipper.SetInsideOut(not invert) 1873 clipper.SetValue(0.0) 1874 1875 clipper.Update() 1876 1877 out = UnstructuredGrid(clipper.GetOutput()) 1878 out.pipeline = utils.OperationNode("cut_with_mesh", parents=[self], c="#9e2a2b") 1879 return out 1880 1881 1882########################################################################## 1883class StructuredGrid(PointAlgorithms, MeshVisual): 1884 """ 1885 Build a structured grid. 1886 """ 1887 1888 def __init__(self, inputobj=None): 1889 """ 1890 A StructuredGrid is a dataset where edges of the hexahedrons are 1891 not necessarily parallel to the coordinate axes. 1892 It can be thought of as a tessellation of a block of 3D space, 1893 similar to a `RectilinearGrid` 1894 except that the cells are not necessarily cubes, they can have different 1895 orientations but are connected in the same way as a `RectilinearGrid`. 1896 1897 Arguments: 1898 inputobj : (vtkStructuredGrid, list, str) 1899 list of points and indices, or filename 1900 1901 Example: 1902 ```python 1903 from vedo import * 1904 sgrid = StructuredGrid(dataurl+"structgrid.vts") 1905 print(sgrid) 1906 msh = sgrid.tomesh().lw(1) 1907 show(msh, axes=1, viewup="z") 1908 ``` 1909 1910 ```python 1911 from vedo import * 1912 1913 cx = np.sqrt(np.linspace(100, 400, 10)) 1914 cy = np.linspace(30, 40, 20) 1915 cz = np.linspace(40, 50, 30) 1916 x, y, z = np.meshgrid(cx, cy, cz) 1917 1918 sgrid1 = StructuredGrid([x, y, z]) 1919 sgrid1.cmap("viridis", sgrid1.coordinates[:, 0]) 1920 print(sgrid1) 1921 1922 sgrid2 = sgrid1.clone().cut_with_plane(normal=(-1,1,1), origin=[14,34,44]) 1923 msh2 = sgrid2.tomesh(shrink=0.9).lw(1).cmap("viridis") 1924 1925 show( 1926 [["StructuredGrid", sgrid1], ["Shrinked Mesh", msh2]], 1927 N=2, axes=1, viewup="z", 1928 ) 1929 ``` 1930 """ 1931 1932 super().__init__() 1933 1934 self.dataset = None 1935 1936 self.mapper = vtki.new("PolyDataMapper") 1937 self._actor = vtki.vtkActor() 1938 self._actor.retrieve_object = weak_ref_to(self) 1939 self._actor.SetMapper(self.mapper) 1940 self.properties = self._actor.GetProperty() 1941 1942 self.transform = LinearTransform() 1943 self.point_locator = None 1944 self.cell_locator = None 1945 self.line_locator = None 1946 1947 self.name = "StructuredGrid" 1948 self.filename = "" 1949 1950 self.info = {} 1951 self.time = time.time() 1952 1953 ############################### 1954 if inputobj is None: 1955 self.dataset = vtki.vtkStructuredGrid() 1956 1957 elif isinstance(inputobj, vtki.vtkStructuredGrid): 1958 self.dataset = inputobj 1959 1960 elif isinstance(inputobj, StructuredGrid): 1961 self.dataset = inputobj.dataset 1962 1963 elif isinstance(inputobj, str): 1964 if "https://" in inputobj: 1965 inputobj = download(inputobj, verbose=False) 1966 if inputobj.endswith(".vts"): 1967 reader = vtki.new("XMLStructuredGridReader") 1968 else: 1969 reader = vtki.new("StructuredGridReader") 1970 self.filename = inputobj 1971 reader.SetFileName(inputobj) 1972 reader.Update() 1973 self.dataset = reader.GetOutput() 1974 1975 elif utils.is_sequence(inputobj): 1976 self.dataset = vtki.vtkStructuredGrid() 1977 x, y, z = inputobj 1978 xyz = np.vstack(( 1979 x.flatten(order="F"), 1980 y.flatten(order="F"), 1981 z.flatten(order="F")) 1982 ).T 1983 dims = x.shape 1984 self.dataset.SetDimensions(dims) 1985 # self.dataset.SetDimensions(dims[1], dims[0], dims[2]) 1986 vpoints = vtki.vtkPoints() 1987 vpoints.SetData(utils.numpy2vtk(xyz)) 1988 self.dataset.SetPoints(vpoints) 1989 1990 1991 ############################### 1992 if not self.dataset: 1993 vedo.logger.error(f"StructuredGrid: cannot understand input type {type(inputobj)}") 1994 return 1995 1996 self.properties.SetColor(0.352, 0.612, 0.996) # blue7 1997 1998 self.pipeline = utils.OperationNode( 1999 self, comment=f"#tets {self.dataset.GetNumberOfCells()}", c="#9e2a2b" 2000 ) 2001 2002 @property 2003 def actor(self): 2004 """Return the `vtkActor` of the object.""" 2005 gf = vtki.new("GeometryFilter") 2006 gf.SetInputData(self.dataset) 2007 gf.Update() 2008 self.mapper.SetInputData(gf.GetOutput()) 2009 self.mapper.Modified() 2010 return self._actor 2011 2012 @actor.setter 2013 def actor(self, _): 2014 pass 2015 2016 def _update(self, data, reset_locators=False): 2017 self.dataset = data 2018 if reset_locators: 2019 self.cell_locator = None 2020 self.point_locator = None 2021 return self 2022 2023 ################################################################## 2024 def __str__(self): 2025 """Print a summary for the `StructuredGrid` object.""" 2026 module = self.__class__.__module__ 2027 name = self.__class__.__name__ 2028 out = vedo.printc( 2029 f"{module}.{name} at ({hex(self.memory_address())})".ljust(75), 2030 c="c", bold=True, invert=True, return_string=True, 2031 ) 2032 out += "\x1b[0m\x1b[36;1m" 2033 2034 out += "name".ljust(14) + ": " + str(self.name) + "\n" 2035 if self.filename: 2036 out += "filename".ljust(14) + ": " + str(self.filename) + "\n" 2037 2038 out += "dimensions".ljust(14) + ": " + str(self.dataset.GetDimensions()) + "\n" 2039 2040 out += "center".ljust(14) + ": " 2041 out += utils.precision(self.dataset.GetCenter(), 6) + "\n" 2042 2043 bnds = self.bounds() 2044 bx1, bx2 = utils.precision(bnds[0], 3), utils.precision(bnds[1], 3) 2045 by1, by2 = utils.precision(bnds[2], 3), utils.precision(bnds[3], 3) 2046 bz1, bz2 = utils.precision(bnds[4], 3), utils.precision(bnds[5], 3) 2047 out += "bounds".ljust(14) + ":" 2048 out += " x=(" + bx1 + ", " + bx2 + ")," 2049 out += " y=(" + by1 + ", " + by2 + ")," 2050 out += " z=(" + bz1 + ", " + bz2 + ")\n" 2051 2052 out += "memory size".ljust(14) + ": " 2053 out += utils.precision(self.dataset.GetActualMemorySize() / 1024, 2) + " MB\n" 2054 2055 for key in self.pointdata.keys(): 2056 arr = self.pointdata[key] 2057 rng = utils.precision(arr.min(), 3) + ", " + utils.precision(arr.max(), 3) 2058 mark_active = "pointdata" 2059 a_scalars = self.dataset.GetPointData().GetScalars() 2060 a_vectors = self.dataset.GetPointData().GetVectors() 2061 a_tensors = self.dataset.GetPointData().GetTensors() 2062 if a_scalars and a_scalars.GetName() == key: 2063 mark_active += " *" 2064 elif a_vectors and a_vectors.GetName() == key: 2065 mark_active += " **" 2066 elif a_tensors and a_tensors.GetName() == key: 2067 mark_active += " ***" 2068 out += mark_active.ljust(14) + f': "{key}" ({arr.dtype}), ndim={arr.ndim}' 2069 out += f", range=({rng})\n" 2070 2071 for key in self.celldata.keys(): 2072 arr = self.celldata[key] 2073 rng = utils.precision(arr.min(), 3) + ", " + utils.precision(arr.max(), 3) 2074 mark_active = "celldata" 2075 a_scalars = self.dataset.GetCellData().GetScalars() 2076 a_vectors = self.dataset.GetCellData().GetVectors() 2077 a_tensors = self.dataset.GetCellData().GetTensors() 2078 if a_scalars and a_scalars.GetName() == key: 2079 mark_active += " *" 2080 elif a_vectors and a_vectors.GetName() == key: 2081 mark_active += " **" 2082 elif a_tensors and a_tensors.GetName() == key: 2083 mark_active += " ***" 2084 out += mark_active.ljust(14) + f': "{key}" ({arr.dtype}), ndim={arr.ndim}' 2085 out += f", range=({rng})\n" 2086 2087 for key in self.metadata.keys(): 2088 arr = self.metadata[key] 2089 out += "metadata".ljust(14) + ": " + f'"{key}" ({len(arr)} values)\n' 2090 2091 return out.rstrip() + "\x1b[0m" 2092 2093 def _repr_html_(self): 2094 """ 2095 HTML representation of the StructuredGrid object for Jupyter Notebooks. 2096 2097 Returns: 2098 HTML text with the image and some properties. 2099 """ 2100 import io 2101 import base64 2102 from PIL import Image 2103 2104 library_name = "vedo.grids.StructuredGrid" 2105 help_url = "https://vedo.embl.es/docs/vedo/grids.html#StructuredGrid" 2106 2107 m = self.tomesh().linewidth(1).lighting("off") 2108 arr= m.thumbnail(zoom=1, elevation=-30, azimuth=-30) 2109 2110 im = Image.fromarray(arr) 2111 buffered = io.BytesIO() 2112 im.save(buffered, format="PNG", quality=100) 2113 encoded = base64.b64encode(buffered.getvalue()).decode("utf-8") 2114 url = "data:image/png;base64," + encoded 2115 image = f"<img src='{url}'></img>" 2116 2117 bounds = "<br/>".join( 2118 [ 2119 utils.precision(min_x, 4) + " ... " + utils.precision(max_x, 4) 2120 for min_x, max_x in zip(self.bounds()[::2], self.bounds()[1::2]) 2121 ] 2122 ) 2123 2124 help_text = "" 2125 if self.name: 2126 help_text += f"<b> {self.name}:   </b>" 2127 help_text += '<b><a href="' + help_url + '" target="_blank">' + library_name + "</a></b>" 2128 if self.filename: 2129 dots = "" 2130 if len(self.filename) > 30: 2131 dots = "..." 2132 help_text += f"<br/><code><i>({dots}{self.filename[-30:]})</i></code>" 2133 2134 pdata = "" 2135 if self.dataset.GetPointData().GetScalars(): 2136 if self.dataset.GetPointData().GetScalars().GetName(): 2137 name = self.dataset.GetPointData().GetScalars().GetName() 2138 pdata = "<tr><td><b> point data array </b></td><td>" + name + "</td></tr>" 2139 2140 cdata = "" 2141 if self.dataset.GetCellData().GetScalars(): 2142 if self.dataset.GetCellData().GetScalars().GetName(): 2143 name = self.dataset.GetCellData().GetScalars().GetName() 2144 cdata = "<tr><td><b> cell data array </b></td><td>" + name + "</td></tr>" 2145 2146 pts = self.coordinates 2147 cm = np.mean(pts, axis=0) 2148 2149 _all = [ 2150 "<table>", 2151 "<tr>", 2152 "<td>", image, "</td>", 2153 "<td style='text-align: center; vertical-align: center;'><br/>", help_text, 2154 "<table>", 2155 "<tr><td><b> bounds </b> <br/> (x/y/z) </td><td>" + str(bounds) + "</td></tr>", 2156 "<tr><td><b> center of mass </b></td><td>" + utils.precision(cm,3) + "</td></tr>", 2157 "<tr><td><b> nr. points / cells </b></td><td>" 2158 + str(self.npoints) + " / " + str(self.ncells) + "</td></tr>", 2159 pdata, 2160 cdata, 2161 "</table>", 2162 "</table>", 2163 ] 2164 return "\n".join(_all) 2165 2166 def dimensions(self) -> np.ndarray: 2167 """Return the number of points in the x, y and z directions.""" 2168 return np.array(self.dataset.GetDimensions()) 2169 2170 def clone(self, deep=True) -> "StructuredGrid": 2171 """Return a clone copy of the StructuredGrid. Alias of `copy()`.""" 2172 if deep: 2173 newrg = vtki.vtkStructuredGrid() 2174 newrg.CopyStructure(self.dataset) 2175 newrg.CopyAttributes(self.dataset) 2176 newvol = StructuredGrid(newrg) 2177 else: 2178 newvol = StructuredGrid(self.dataset) 2179 2180 prop = vtki.vtkProperty() 2181 prop.DeepCopy(self.properties) 2182 newvol.actor.SetProperty(prop) 2183 newvol.properties = prop 2184 newvol.pipeline = utils.OperationNode("clone", parents=[self], c="#bbd0ff", shape="diamond") 2185 return newvol 2186 2187 def find_point(self, x: list) -> int: 2188 """Given a position `x`, return the id of the closest point.""" 2189 return self.dataset.FindPoint(x) 2190 2191 def find_cell(self, x: list) -> dict: 2192 """Given a position `x`, return the id of the closest cell.""" 2193 cell = vtki.vtkHexagonalPrism() 2194 cellid = vtki.mutable(0) 2195 tol2 = 0.001 # vtki.mutable(0) 2196 subid = vtki.mutable(0) 2197 pcoords = [0.0, 0.0, 0.0] 2198 weights = [0.0, 0.0, 0.0] 2199 res = self.dataset.FindCell(x, cell, cellid, tol2, subid, pcoords, weights) 2200 result = {} 2201 result["cellid"] = cellid 2202 result["subid"] = subid 2203 result["pcoords"] = pcoords 2204 result["weights"] = weights 2205 result["status"] = res 2206 return result 2207 2208 def cut_with_plane(self, origin=(0, 0, 0), normal="x") -> "vedo.UnstructuredGrid": 2209 """ 2210 Cut the object with the plane defined by a point and a normal. 2211 2212 Arguments: 2213 origin : (list) 2214 the cutting plane goes through this point 2215 normal : (list, str) 2216 normal vector to the cutting plane 2217 2218 Returns an `UnstructuredGrid` object. 2219 """ 2220 strn = str(normal) 2221 if strn == "x": normal = (1, 0, 0) 2222 elif strn == "y": normal = (0, 1, 0) 2223 elif strn == "z": normal = (0, 0, 1) 2224 elif strn == "-x": normal = (-1, 0, 0) 2225 elif strn == "-y": normal = (0, -1, 0) 2226 elif strn == "-z": normal = (0, 0, -1) 2227 plane = vtki.new("Plane") 2228 plane.SetOrigin(origin) 2229 plane.SetNormal(normal) 2230 clipper = vtki.new("ClipDataSet") 2231 clipper.SetInputData(self.dataset) 2232 clipper.SetClipFunction(plane) 2233 clipper.GenerateClipScalarsOff() 2234 clipper.GenerateClippedOutputOff() 2235 clipper.SetValue(0) 2236 clipper.Update() 2237 cout = clipper.GetOutput() 2238 ug = vedo.UnstructuredGrid(cout) 2239 if isinstance(self, vedo.UnstructuredGrid): 2240 self._update(cout) 2241 self.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 2242 return self 2243 ug.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 2244 return ug 2245 2246 def cut_with_mesh(self, mesh: Mesh, invert=False, whole_cells=False, on_boundary=False) -> "UnstructuredGrid": 2247 """ 2248 Cut a `RectilinearGrid` with a `Mesh`. 2249 2250 Use `invert` to return cut off part of the input object. 2251 2252 Returns an `UnstructuredGrid` object. 2253 """ 2254 ug = self.dataset 2255 2256 ippd = vtki.new("ImplicitPolyDataDistance") 2257 ippd.SetInput(mesh.dataset) 2258 2259 if whole_cells or on_boundary: 2260 clipper = vtki.new("ExtractGeometry") 2261 clipper.SetInputData(ug) 2262 clipper.SetImplicitFunction(ippd) 2263 clipper.SetExtractInside(not invert) 2264 clipper.SetExtractBoundaryCells(False) 2265 if on_boundary: 2266 clipper.SetExtractBoundaryCells(True) 2267 clipper.SetExtractOnlyBoundaryCells(True) 2268 else: 2269 signed_dists = vtki.vtkFloatArray() 2270 signed_dists.SetNumberOfComponents(1) 2271 signed_dists.SetName("SignedDistance") 2272 for pointId in range(ug.GetNumberOfPoints()): 2273 p = ug.GetPoint(pointId) 2274 signed_dist = ippd.EvaluateFunction(p) 2275 signed_dists.InsertNextValue(signed_dist) 2276 ug.GetPointData().AddArray(signed_dists) 2277 ug.GetPointData().SetActiveScalars("SignedDistance") # NEEDED 2278 clipper = vtki.new("ClipDataSet") 2279 clipper.SetInputData(ug) 2280 clipper.SetInsideOut(not invert) 2281 clipper.SetValue(0.0) 2282 2283 clipper.Update() 2284 2285 out = UnstructuredGrid(clipper.GetOutput()) 2286 out.pipeline = utils.OperationNode("cut_with_mesh", parents=[self], c="#9e2a2b") 2287 return out 2288 2289 def isosurface(self, value=None) -> "vedo.Mesh": 2290 """ 2291 Return a `Mesh` isosurface extracted from the object. 2292 2293 Set `value` as single float or list of values to draw the isosurface(s). 2294 """ 2295 scrange = self.dataset.GetScalarRange() 2296 2297 cf = vtki.new("ContourFilter") 2298 cf.UseScalarTreeOn() 2299 cf.SetInputData(self.dataset) 2300 cf.ComputeNormalsOn() 2301 2302 if value is None: 2303 value = (2 * scrange[0] + scrange[1]) / 3.0 2304 # print("automatic isosurface value =", value) 2305 cf.SetValue(0, value) 2306 else: 2307 if utils.is_sequence(value): 2308 cf.SetNumberOfContours(len(value)) 2309 for i, t in enumerate(value): 2310 cf.SetValue(i, t) 2311 else: 2312 cf.SetValue(0, value) 2313 2314 cf.Update() 2315 poly = cf.GetOutput() 2316 2317 out = vedo.mesh.Mesh(poly, c=None).phong() 2318 out.mapper.SetScalarRange(scrange[0], scrange[1]) 2319 2320 out.pipeline = utils.OperationNode( 2321 "isosurface", 2322 parents=[self], 2323 comment=f"#pts {out.dataset.GetNumberOfPoints()}", 2324 c="#4cc9f0:#e9c46a", 2325 ) 2326 return out 2327 2328 2329 2330######### WIP ############## 2331class ExplicitStructuredGrid: 2332 """ 2333 Build an explicit structured grid. 2334 2335 An explicit structured grid is a dataset where edges of the hexahedrons are 2336 not necessarily parallel to the coordinate axes. 2337 It can be thought of as a tessellation of a block of 3D space, 2338 similar to a `RectilinearGrid` 2339 except that the cells are not necessarily cubes, they can have different 2340 orientations but are connected in the same way as a `RectilinearGrid`. 2341 2342 Arguments: 2343 inputobj : (vtkExplicitStructuredGrid, list, str) 2344 list of points and indices, or filename 2345 """ 2346 2347 # int GetDataDimension () 2348 # Return the dimensionality of the data. 2349 2350 # void GetCellDims (int cellDims[3]) 2351 # Computes the cell dimensions according to internal point dimensions. 2352 2353 # int GetExtentType () override 2354 # The extent type is a 3D extent. 2355 2356 # void BuildLinks () 2357 # Build topological links from points to lists of cells that use each point. 2358 2359 # vtkIdType * GetCellPoints (vtkIdType cellId) 2360 # Get direct raw pointer to the 8 points indices of an hexahedra. 2361 2362 # void GetCellPoints (vtkIdType cellId, vtkIdType &npts, vtkIdType *&pts) 2363 # More efficient method to obtain cell points. 2364 2365 # void GetCellPoints (vtkIdType cellId, vtkIdType &npts, vtkIdType const *&pts, vtkIdList *ptIds) override 2366 # More efficient method to obtain cell points. 2367 2368 # void GetCellNeighbors (vtkIdType cellId, vtkIdType neighbors[6], int *wholeExtent=nullptr) 2369 # Get cell neighbors of the cell for every faces. 2370 2371 # void ComputeCellStructuredCoords (vtkIdType cellId, int &i, int &j, int &k, bool adjustForExtent=true) 2372 # Given a cellId, get the structured coordinates (i-j-k). 2373 2374 # vtkIdType ComputeCellId (int i, int j, int k, bool adjustForExtent=true) 2375 # Given a location in structured coordinates (i-j-k), return the cell id. 2376 2377 # void ComputeFacesConnectivityFlagsArray () 2378 # Compute the faces connectivity flags array. 2379 2380 # bool HasAnyBlankCells () override 2381 # Returns true if one or more cells are blanked, false otherwise. 2382 2383 # unsigned char IsCellVisible (vtkIdType cellId) 2384 # Return non-zero value if specified cell is visible. 2385 2386 # unsigned char IsCellGhost (vtkIdType cellId) 2387 # Return non-zero value if specified cell is a ghost cell. 2388 2389 # bool HasAnyGhostCells () 2390 # Returns true if one or more cells are ghost, false otherwise. 2391 2392 # void CheckAndReorderFaces () 2393 # Check faces are numbered correctly regarding ijk numbering 2394 # If not this will reorganize cell points order so face order is valid. 2395 2396 # void GetCellBounds (vtkIdType cellId, double bounds[6]) override 2397 # Standard vtkDataSet API methods. 2398 2399 # int GetCellType (vtkIdType cellId) override 2400 # Standard vtkDataSet API methods. 2401 2402 # vtkIdType GetCellSize (vtkIdType cellId) override 2403 # Standard vtkDataSet API methods. 2404 2405 # vtkIdType GetNumberOfCells () override 2406 # Standard vtkDataSet API methods. 2407 2408 # void GetCellPoints (vtkIdType cellId, vtkIdList *ptIds) override 2409 # Standard vtkDataSet API methods. 2410 2411 # void GetPointCells (vtkIdType ptId, vtkIdList *cellIds) override 2412 # Standard vtkDataSet API methods. 2413 2414 # int GetMaxCellSize () override 2415 # Standard vtkDataSet API methods. 2416 2417 # int GetMaxSpatialDimension () override 2418 # Standard vtkDataSet API methods. 2419 2420 # int GetMinSpatialDimension () override 2421 # Standard vtkDataSet API methods. 2422 2423 # void GetCellNeighbors (vtkIdType cellId, vtkIdList *ptIds, vtkIdList *cellIds) override 2424 # Standard vtkDataSet API methods. 2425 2426 # void SetDimensions (int i, int j, int k) 2427 # Set/Get the dimensions of this structured dataset in term of number of points along each direction. 2428 2429 # void SetDimensions (int dim[3]) 2430 # Set/Get the dimensions of this structured dataset in term of number of points along each direction. 2431 2432 # void GetDimensions (int dim[3]) 2433 # Set/Get the dimensions of this structured dataset in term of number of points along each direction. 2434 2435 # void SetExtent (int x0, int x1, int y0, int y1, int z0, int z1) 2436 # Set/Get the extent of this structured dataset in term of number of points along each direction. 2437 2438 # void SetExtent (int extent[6]) 2439 # Set/Get the extent of this structured dataset in term of number of points along each direction. 2440 2441 def __init__(self, inputobj=None): 2442 """ 2443 A StructuredGrid is a dataset where edges of the hexahedrons are 2444 not necessarily parallel to the coordinate axes. 2445 It can be thought of as a tessellation of a block of 3D space, 2446 similar to a `RectilinearGrid` 2447 except that the cells are not necessarily cubes, they can have different 2448 orientations but are connected in the same way as a `RectilinearGrid`. 2449 2450 Arguments: 2451 inputobj : (vtkExplicitStructuredGrid, list, str) 2452 list of points and indices, or filename" 2453 """ 2454 self.dataset = None 2455 self.mapper = vtki.new("PolyDataMapper") 2456 self._actor = vtki.vtkActor() 2457 self._actor.retrieve_object = weak_ref_to(self) 2458 self._actor.SetMapper(self.mapper) 2459 self.properties = self._actor.GetProperty() 2460 2461 self.transform = LinearTransform() 2462 self.point_locator = None 2463 self.cell_locator = None 2464 self.line_locator = None 2465 2466 self.name = "ExplicitStructuredGrid" 2467 self.filename = "" 2468 2469 self.info = {} 2470 self.time = time.time() 2471 2472 ############################### 2473 if inputobj is None: 2474 self.dataset = vtki.vtkExplicitStructuredGrid() 2475 2476 elif isinstance(inputobj, vtki.vtkExplicitStructuredGrid): 2477 self.dataset = inputobj 2478 2479 elif isinstance(inputobj, ExplicitStructuredGrid): 2480 self.dataset = inputobj.dataset 2481 2482 elif isinstance(inputobj, str): 2483 if "https://" in inputobj: 2484 inputobj = download(inputobj, verbose=False) 2485 if inputobj.endswith(".vts"): 2486 reader = vtki.new("XMLExplicitStructuredGridReader") 2487 else: 2488 reader = vtki.new("ExplicitStructuredGridReader") 2489 self.filename = inputobj 2490 reader.SetFileName(inputobj) 2491 reader.Update() 2492 self.dataset = reader.GetOutput() 2493 2494 elif utils.is_sequence(inputobj): 2495 self.dataset = vtki.vtkExplicitStructuredGrid() 2496 x, y, z = inputobj 2497 xyz = np.vstack(( 2498 x.flatten(order="F"), 2499 y.flatten(order="F"), 2500 z.flatten(order="F")) 2501 ).T 2502 dims = x.shape 2503 self.dataset.SetDimensions(dims) 2504 # self.dataset.SetDimensions(dims[1], dims[0], dims[2]) 2505 vpoints = vtki.vtkPoints() 2506 vpoints.SetData(utils.numpy2vtk(xyz)) 2507 self.dataset.SetPoints(vpoints) 2508 2509 2510 ############################### 2511 if not self.dataset: 2512 vedo.logger.error(f"ExplicitStructuredGrid: cannot understand input type {type(inputobj)}") 2513 return 2514 2515 self.properties.SetColor(0.352, 0.612, 0.996) # blue7 2516 self.pipeline = utils.OperationNode( 2517 self, comment=f"#cells {self.dataset.GetNumberOfCells()}", c="#9e2a2b" 2518 ) 2519 2520 @property 2521 def actor(self): 2522 """Return the `vtkActor` of the object.""" 2523 gf = vtki.new("GeometryFilter") 2524 gf.SetInputData(self.dataset) 2525 gf.Update() 2526 self.mapper.SetInputData(gf.GetOutput()) 2527 self.mapper.Modified() 2528 return self._actor 2529 2530 @actor.setter 2531 def actor(self, _): 2532 pass 2533 2534 def _update(self, data, reset_locators=False): 2535 self.dataset = data 2536 if reset_locators: 2537 self.cell_locator = None 2538 self.point_locator = None 2539 return self 2540 2541 def dimensions(self) -> np.ndarray: 2542 """Return the number of points in the x, y and z directions.""" 2543 return np.array(self.dataset.GetDimensions()) 2544 2545 def clone(self, deep=True) -> "ExplicitStructuredGrid": 2546 """Return a clone copy of the StructuredGrid. Alias of `copy()`.""" 2547 if deep: 2548 newrg = vtki.vtkExplicitStructuredGrid() 2549 newrg.CopyStructure(self.dataset) 2550 newrg.CopyAttributes(self.dataset) 2551 newvol = ExplicitStructuredGrid(newrg) 2552 else: 2553 newvol = ExplicitStructuredGrid(self.dataset) 2554 2555 prop = vtki.vtkProperty() 2556 prop.DeepCopy(self.properties) 2557 newvol.actor.SetProperty(prop) 2558 newvol.properties = prop 2559 newvol.pipeline = utils.OperationNode("clone", parents=[self], c="#bbd0ff", shape="diamond") 2560 return newvol 2561 2562 def cut_with_plane(self, origin=(0, 0, 0), normal="x") -> "vedo.UnstructuredGrid": 2563 """ 2564 Cut the object with the plane defined by a point and a normal. 2565 2566 Arguments: 2567 origin : (list) 2568 the cutting plane goes through this point 2569 normal : (list, str) 2570 normal vector to the cutting plane 2571 2572 Returns an `UnstructuredGrid` object. 2573 """ 2574 strn = str(normal) 2575 if strn == "x": normal = (1, 0, 0) 2576 elif strn == "y": normal = (0, 1, 0) 2577 elif strn == "z": normal = (0, 0, 1) 2578 elif strn == "-x": normal = (-1, 0, 0) 2579 elif strn == "-y": normal = (0, -1, 0) 2580 elif strn == "-z": normal = (0, 0, -1) 2581 plane = vtki.new("Plane") 2582 plane.SetOrigin(origin) 2583 plane.SetNormal(normal) 2584 clipper = vtki.new("ClipDataSet") 2585 clipper.SetInputData(self.dataset) 2586 clipper.SetClipFunction(plane) 2587 clipper.GenerateClipScalarsOff() 2588 clipper.GenerateClippedOutputOff() 2589 clipper.SetValue(0) 2590 clipper.Update() 2591 cout = clipper.GetOutput() 2592 ug = vedo.UnstructuredGrid(cout) 2593 if isinstance(self, vedo.UnstructuredGrid): 2594 self._update(cout) 2595 self.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 2596 return self 2597 ug.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 2598 return ug 2599
38class UnstructuredGrid(PointAlgorithms, MeshVisual): 39 """Support for UnstructuredGrid objects.""" 40 41 def __init__(self, inputobj=None): 42 """ 43 Support for UnstructuredGrid objects. 44 45 Arguments: 46 inputobj : (list, vtkUnstructuredGrid, str) 47 A list in the form `[points, cells, celltypes]`, 48 or a vtkUnstructuredGrid object, or a filename 49 50 Celltypes are identified by the following 51 [convention](https://vtk.org/doc/nightly/html/vtkCellType_8h_source.html). 52 """ 53 super().__init__() 54 55 self.dataset = None 56 57 self.mapper = vtki.new("PolyDataMapper") 58 self._actor = vtki.vtkActor() 59 self._actor.retrieve_object = weak_ref_to(self) 60 self._actor.SetMapper(self.mapper) 61 self.properties = self._actor.GetProperty() 62 63 self.transform = LinearTransform() 64 self.point_locator = None 65 self.cell_locator = None 66 self.line_locator = None 67 68 self.name = "UnstructuredGrid" 69 self.filename = "" 70 self.file_size = "" 71 72 self.info = {} 73 self.time = time.time() 74 self.rendered_at = set() 75 76 ###################################### 77 inputtype = str(type(inputobj)) 78 79 if inputobj is None: 80 self.dataset = vtki.vtkUnstructuredGrid() 81 82 elif utils.is_sequence(inputobj): 83 84 pts, cells, celltypes = inputobj 85 assert len(cells) == len(celltypes) 86 87 self.dataset = vtki.vtkUnstructuredGrid() 88 89 if not utils.is_sequence(cells[0]): 90 tets = [] 91 nf = cells[0] + 1 92 for i, cl in enumerate(cells): 93 if i in (nf, 0): 94 k = i + 1 95 nf = cl + k 96 cell = [cells[j + k] for j in range(cl)] 97 tets.append(cell) 98 cells = tets 99 100 # This would fill the points and use those to define orientation 101 vpts = utils.numpy2vtk(pts, dtype=np.float32) 102 points = vtki.vtkPoints() 103 points.SetData(vpts) 104 self.dataset.SetPoints(points) 105 106 # Fill cells 107 # https://vtk.org/doc/nightly/html/vtkCellType_8h_source.html 108 for i, ct in enumerate(celltypes): 109 if ct == vtki.cell_types["VERTEX"]: 110 cell = vtki.vtkVertex() 111 elif ct == vtki.cell_types["POLY_VERTEX"]: 112 cell = vtki.vtkPolyVertex() 113 elif ct == vtki.cell_types["TETRA"]: 114 cell = vtki.vtkTetra() 115 elif ct == vtki.cell_types["WEDGE"]: 116 cell = vtki.vtkWedge() 117 elif ct == vtki.cell_types["LINE"]: 118 cell = vtki.vtkLine() 119 elif ct == vtki.cell_types["POLY_LINE"]: 120 cell = vtki.vtkPolyLine() 121 elif ct == vtki.cell_types["TRIANGLE"]: 122 cell = vtki.vtkTriangle() 123 elif ct == vtki.cell_types["TRIANGLE_STRIP"]: 124 cell = vtki.vtkTriangleStrip() 125 elif ct == vtki.cell_types["POLYGON"]: 126 cell = vtki.vtkPolygon() 127 elif ct == vtki.cell_types["PIXEL"]: 128 cell = vtki.vtkPixel() 129 elif ct == vtki.cell_types["QUAD"]: 130 cell = vtki.vtkQuad() 131 elif ct == vtki.cell_types["VOXEL"]: 132 cell = vtki.vtkVoxel() 133 elif ct == vtki.cell_types["PYRAMID"]: 134 cell = vtki.vtkPyramid() 135 elif ct == vtki.cell_types["HEXAHEDRON"]: 136 cell = vtki.vtkHexahedron() 137 elif ct == vtki.cell_types["HEXAGONAL_PRISM"]: 138 cell = vtki.vtkHexagonalPrism() 139 elif ct == vtki.cell_types["PENTAGONAL_PRISM"]: 140 cell = vtki.vtkPentagonalPrism() 141 elif ct == vtki.cell_types["QUADRATIC_TETRA"]: 142 from vtkmodules.vtkCommonDataModel import vtkQuadraticTetra 143 cell = vtkQuadraticTetra() 144 elif ct == vtki.cell_types["QUADRATIC_HEXAHEDRON"]: 145 from vtkmodules.vtkCommonDataModel import vtkQuadraticHexahedron 146 cell = vtkQuadraticHexahedron() 147 elif ct == vtki.cell_types["QUADRATIC_WEDGE"]: 148 from vtkmodules.vtkCommonDataModel import vtkQuadraticWedge 149 cell = vtkQuadraticWedge() 150 elif ct == vtki.cell_types["QUADRATIC_PYRAMID"]: 151 from vtkmodules.vtkCommonDataModel import vtkQuadraticPyramid 152 cell = vtkQuadraticPyramid() 153 elif ct == vtki.cell_types["QUADRATIC_LINEAR_QUAD"]: 154 from vtkmodules.vtkCommonDataModel import vtkQuadraticLinearQuad 155 cell = vtkQuadraticLinearQuad() 156 elif ct == vtki.cell_types["QUADRATIC_LINEAR_WEDGE"]: 157 from vtkmodules.vtkCommonDataModel import vtkQuadraticLinearWedge 158 cell = vtkQuadraticLinearWedge() 159 elif ct == vtki.cell_types["BIQUADRATIC_QUADRATIC_WEDGE"]: 160 from vtkmodules.vtkCommonDataModel import vtkBiQuadraticQuadraticWedge 161 cell = vtkBiQuadraticQuadraticWedge() 162 elif ct == vtki.cell_types["BIQUADRATIC_QUADRATIC_HEXAHEDRON"]: 163 from vtkmodules.vtkCommonDataModel import vtkBiQuadraticQuadraticHexahedron 164 cell = vtkBiQuadraticQuadraticHexahedron() 165 elif ct == vtki.cell_types["BIQUADRATIC_TRIANGLE"]: 166 from vtkmodules.vtkCommonDataModel import vtkBiQuadraticTriangle 167 cell = vtkBiQuadraticTriangle() 168 elif ct == vtki.cell_types["CUBIC_LINE"]: 169 from vtkmodules.vtkCommonDataModel import vtkCubicLine 170 cell = vtkCubicLine() 171 elif ct == vtki.cell_types["CONVEX_POINT_SET"]: 172 from vtkmodules.vtkCommonDataModel import vtkConvexPointSet 173 cell = vtkConvexPointSet() 174 elif ct == vtki.cell_types["POLYHEDRON"]: 175 from vtkmodules.vtkCommonDataModel import vtkPolyhedron 176 cell = vtkPolyhedron() 177 elif ct == vtki.cell_types["HIGHER_ORDER_TRIANGLE"]: 178 from vtkmodules.vtkCommonDataModel import vtkHigherOrderTriangle 179 cell = vtkHigherOrderTriangle() 180 elif ct == vtki.cell_types["HIGHER_ORDER_QUAD"]: 181 from vtkmodules.vtkCommonDataModel import vtkHigherOrderQuadrilateral 182 cell = vtkHigherOrderQuadrilateral() 183 elif ct == vtki.cell_types["HIGHER_ORDER_TETRAHEDRON"]: 184 from vtkmodules.vtkCommonDataModel import vtkHigherOrderTetra 185 cell = vtkHigherOrderTetra() 186 elif ct == vtki.cell_types["HIGHER_ORDER_WEDGE"]: 187 from vtkmodules.vtkCommonDataModel import vtkHigherOrderWedge 188 cell = vtkHigherOrderWedge() 189 elif ct == vtki.cell_types["HIGHER_ORDER_HEXAHEDRON"]: 190 from vtkmodules.vtkCommonDataModel import vtkHigherOrderHexahedron 191 cell = vtkHigherOrderHexahedron() 192 elif ct == vtki.cell_types["LAGRANGE_CURVE"]: 193 from vtkmodules.vtkCommonDataModel import vtkLagrangeCurve 194 cell = vtkLagrangeCurve() 195 elif ct == vtki.cell_types["LAGRANGE_TRIANGLE"]: 196 from vtkmodules.vtkCommonDataModel import vtkLagrangeTriangle 197 cell = vtkLagrangeTriangle() 198 elif ct == vtki.cell_types["LAGRANGE_QUADRILATERAL"]: 199 from vtkmodules.vtkCommonDataModel import vtkLagrangeQuadrilateral 200 cell = vtkLagrangeQuadrilateral() 201 elif ct == vtki.cell_types["LAGRANGE_TETRAHEDRON"]: 202 from vtkmodules.vtkCommonDataModel import vtkLagrangeTetra 203 cell = vtkLagrangeTetra() 204 elif ct == vtki.cell_types["LAGRANGE_HEXAHEDRON"]: 205 from vtkmodules.vtkCommonDataModel import vtkLagrangeHexahedron 206 cell = vtkLagrangeHexahedron() 207 elif ct == vtki.cell_types["LAGRANGE_WEDGE"]: 208 from vtkmodules.vtkCommonDataModel import vtkLagrangeWedge 209 cell = vtkLagrangeWedge() 210 elif ct == vtki.cell_types["BEZIER_CURVE"]: 211 from vtkmodules.vtkCommonDataModel import vtkBezierCurve 212 cell = vtkBezierCurve() 213 elif ct == vtki.cell_types["BEZIER_TRIANGLE"]: 214 from vtkmodules.vtkCommonDataModel import vtkBezierTriangle 215 cell = vtkBezierTriangle() 216 elif ct == vtki.cell_types["BEZIER_QUADRILATERAL"]: 217 from vtkmodules.vtkCommonDataModel import vtkBezierQuadrilateral 218 cell = vtkBezierQuadrilateral() 219 elif ct == vtki.cell_types["BEZIER_TETRAHEDRON"]: 220 from vtkmodules.vtkCommonDataModel import vtkBezierTetra 221 cell = vtkBezierTetra() 222 elif ct == vtki.cell_types["BEZIER_HEXAHEDRON"]: 223 from vtkmodules.vtkCommonDataModel import vtkBezierHexahedron 224 cell = vtkBezierHexahedron() 225 elif ct == vtki.cell_types["BEZIER_WEDGE"]: 226 from vtkmodules.vtkCommonDataModel import vtkBezierWedge 227 cell = vtkBezierWedge() 228 else: 229 vedo.logger.error( 230 f"UnstructuredGrid: cell type {ct} not supported. Skip.") 231 continue 232 233 cpids = cell.GetPointIds() 234 cell_conn = cells[i] 235 for j, pid in enumerate(cell_conn): 236 cpids.SetId(j, pid) 237 self.dataset.InsertNextCell(ct, cpids) 238 239 elif "UnstructuredGrid" in inputtype: 240 self.dataset = inputobj 241 242 elif isinstance(inputobj, str): 243 if "https://" in inputobj: 244 inputobj = download(inputobj, verbose=False) 245 self.filename = inputobj 246 if inputobj.endswith(".vtu"): 247 reader = vtki.new("XMLUnstructuredGridReader") 248 else: 249 reader = vtki.new("UnstructuredGridReader") 250 self.filename = inputobj 251 reader.SetFileName(inputobj) 252 reader.Update() 253 self.dataset = reader.GetOutput() 254 255 else: 256 # this converts other types of vtk objects to UnstructuredGrid 257 apf = vtki.new("AppendFilter") 258 try: 259 apf.AddInputData(inputobj) 260 except TypeError: 261 apf.AddInputData(inputobj.dataset) 262 apf.Update() 263 self.dataset = apf.GetOutput() 264 265 self.properties.SetColor(0.89, 0.455, 0.671) # pink7 266 267 self.pipeline = utils.OperationNode( 268 self, comment=f"#cells {self.dataset.GetNumberOfCells()}", c="#4cc9f0" 269 ) 270 271 # ------------------------------------------------------------------ 272 def __str__(self): 273 """Print a string summary of the `UnstructuredGrid` object.""" 274 module = self.__class__.__module__ 275 name = self.__class__.__name__ 276 out = vedo.printc( 277 f"{module}.{name} at ({hex(self.memory_address())})".ljust(75), 278 c="m", bold=True, invert=True, return_string=True, 279 ) 280 out += "\x1b[0m\u001b[35m" 281 282 out += "nr. of verts".ljust(14) + ": " + str(self.npoints) + "\n" 283 out += "nr. of cells".ljust(14) + ": " + str(self.ncells) + "\n" 284 ct_arr = np.unique(self.cell_types_array) 285 cnames = [k for k, v in vtki.cell_types.items() if v in ct_arr] 286 out += "cell types".ljust(14) + ": " + str(cnames) + "\n" 287 288 if self.npoints: 289 out+="size".ljust(14)+ ": average=" + utils.precision(self.average_size(),6) 290 out+=", diagonal="+ utils.precision(self.diagonal_size(), 6)+ "\n" 291 out+="center of mass".ljust(14) + ": " + utils.precision(self.center_of_mass(),6)+"\n" 292 293 bnds = self.bounds() 294 bx1, bx2 = utils.precision(bnds[0], 3), utils.precision(bnds[1], 3) 295 by1, by2 = utils.precision(bnds[2], 3), utils.precision(bnds[3], 3) 296 bz1, bz2 = utils.precision(bnds[4], 3), utils.precision(bnds[5], 3) 297 out += "bounds".ljust(14) + ":" 298 out += " x=(" + bx1 + ", " + bx2 + ")," 299 out += " y=(" + by1 + ", " + by2 + ")," 300 out += " z=(" + bz1 + ", " + bz2 + ")\n" 301 302 for key in self.pointdata.keys(): 303 arr = self.pointdata[key] 304 rng = utils.precision(arr.min(), 3) + ", " + utils.precision(arr.max(), 3) 305 mark_active = "pointdata" 306 a_scalars = self.dataset.GetPointData().GetScalars() 307 a_vectors = self.dataset.GetPointData().GetVectors() 308 a_tensors = self.dataset.GetPointData().GetTensors() 309 if a_scalars and a_scalars.GetName() == key: 310 mark_active += " *" 311 elif a_vectors and a_vectors.GetName() == key: 312 mark_active += " **" 313 elif a_tensors and a_tensors.GetName() == key: 314 mark_active += " ***" 315 out += mark_active.ljust(14) + f': "{key}" ({arr.dtype}), ndim={arr.ndim}' 316 out += f", range=({rng})\n" 317 318 for key in self.celldata.keys(): 319 arr = self.celldata[key] 320 rng = utils.precision(arr.min(), 3) + ", " + utils.precision(arr.max(), 3) 321 mark_active = "celldata" 322 a_scalars = self.dataset.GetCellData().GetScalars() 323 a_vectors = self.dataset.GetCellData().GetVectors() 324 a_tensors = self.dataset.GetCellData().GetTensors() 325 if a_scalars and a_scalars.GetName() == key: 326 mark_active += " *" 327 elif a_vectors and a_vectors.GetName() == key: 328 mark_active += " **" 329 elif a_tensors and a_tensors.GetName() == key: 330 mark_active += " ***" 331 out += mark_active.ljust(14) + f': "{key}" ({arr.dtype}), ndim={arr.ndim}' 332 out += f", range=({rng})\n" 333 334 for key in self.metadata.keys(): 335 arr = self.metadata[key] 336 out += "metadata".ljust(14) + ": " + f'"{key}" ({len(arr)} values)\n' 337 338 return out.rstrip() + "\x1b[0m" 339 340 def _repr_html_(self): 341 """ 342 HTML representation of the UnstructuredGrid object for Jupyter Notebooks. 343 344 Returns: 345 HTML text with the image and some properties. 346 """ 347 import io 348 import base64 349 from PIL import Image 350 351 library_name = "vedo.grids.UnstructuredGrid" 352 help_url = "https://vedo.embl.es/docs/vedo/grids.html#UnstructuredGrid" 353 354 arr = self.thumbnail() 355 im = Image.fromarray(arr) 356 buffered = io.BytesIO() 357 im.save(buffered, format="PNG", quality=100) 358 encoded = base64.b64encode(buffered.getvalue()).decode("utf-8") 359 url = "data:image/png;base64," + encoded 360 image = f"<img src='{url}'></img>" 361 362 bounds = "<br/>".join( 363 [ 364 utils.precision(min_x, 4) + " ... " + utils.precision(max_x, 4) 365 for min_x, max_x in zip(self.bounds()[::2], self.bounds()[1::2]) 366 ] 367 ) 368 369 help_text = "" 370 if self.name: 371 help_text += f"<b> {self.name}:   </b>" 372 help_text += '<b><a href="' + help_url + '" target="_blank">' + library_name + "</a></b>" 373 if self.filename: 374 dots = "" 375 if len(self.filename) > 30: 376 dots = "..." 377 help_text += f"<br/><code><i>({dots}{self.filename[-30:]})</i></code>" 378 379 pdata = "" 380 if self.dataset.GetPointData().GetScalars(): 381 if self.dataset.GetPointData().GetScalars().GetName(): 382 name = self.dataset.GetPointData().GetScalars().GetName() 383 pdata = "<tr><td><b> point data array </b></td><td>" + name + "</td></tr>" 384 385 cdata = "" 386 if self.dataset.GetCellData().GetScalars(): 387 if self.dataset.GetCellData().GetScalars().GetName(): 388 name = self.dataset.GetCellData().GetScalars().GetName() 389 cdata = "<tr><td><b> cell data array </b></td><td>" + name + "</td></tr>" 390 391 pts = self.coordinates 392 cm = np.mean(pts, axis=0) 393 394 _all = [ 395 "<table>", 396 "<tr>", 397 "<td>", image, "</td>", 398 "<td style='text-align: center; vertical-align: center;'><br/>", help_text, 399 "<table>", 400 "<tr><td><b> bounds </b> <br/> (x/y/z) </td><td>" + str(bounds) + "</td></tr>", 401 "<tr><td><b> center of mass </b></td><td>" + utils.precision(cm,3) + "</td></tr>", 402 # "<tr><td><b> average size </b></td><td>" + str(average_size) + "</td></tr>", 403 "<tr><td><b> nr. points / cells </b></td><td>" 404 + str(self.npoints) + " / " + str(self.ncells) + "</td></tr>", 405 pdata, 406 cdata, 407 "</table>", 408 "</table>", 409 ] 410 return "\n".join(_all) 411 412 @property 413 def actor(self): 414 """Return the `vtkActor` of the object.""" 415 # print("building actor") 416 gf = vtki.new("GeometryFilter") 417 gf.SetInputData(self.dataset) 418 gf.Update() 419 out = gf.GetOutput() 420 self.mapper.SetInputData(out) 421 self.mapper.Modified() 422 return self._actor 423 424 @actor.setter 425 def actor(self, _): 426 pass 427 428 def _update(self, data, reset_locators=False): 429 self.dataset = data 430 if reset_locators: 431 self.cell_locator = None 432 self.point_locator = None 433 return self 434 435 def merge(self, *others) -> Self: 436 """ 437 Merge multiple datasets into one single `UnstrcturedGrid`. 438 """ 439 apf = vtki.new("AppendFilter") 440 for o in others: 441 if isinstance(o, UnstructuredGrid): 442 apf.AddInputData(o.dataset) 443 elif isinstance(o, vtki.vtkUnstructuredGrid): 444 apf.AddInputData(o) 445 else: 446 vedo.printc("Error: cannot merge type", type(o), c="r") 447 apf.Update() 448 self._update(apf.GetOutput()) 449 self.pipeline = utils.OperationNode( 450 "merge", parents=[self, *others], c="#9e2a2b" 451 ) 452 return self 453 454 def copy(self, deep=True) -> "UnstructuredGrid": 455 """Return a copy of the object. Alias of `clone()`.""" 456 return self.clone(deep=deep) 457 458 def clone(self, deep=True) -> "UnstructuredGrid": 459 """Clone the UnstructuredGrid object to yield an exact copy.""" 460 ug = vtki.vtkUnstructuredGrid() 461 if deep: 462 ug.DeepCopy(self.dataset) 463 else: 464 ug.ShallowCopy(self.dataset) 465 if isinstance(self, vedo.UnstructuredGrid): 466 cloned = vedo.UnstructuredGrid(ug) 467 else: 468 cloned = vedo.TetMesh(ug) 469 470 cloned.copy_properties_from(self) 471 472 cloned.pipeline = utils.OperationNode( 473 "clone", parents=[self], shape="diamond", c="#bbe1ed" 474 ) 475 return cloned 476 477 def bounds(self) -> np.ndarray: 478 """ 479 Get the object bounds. 480 Returns a list in format `[xmin,xmax, ymin,ymax, zmin,zmax]`. 481 """ 482 # OVERRIDE CommonAlgorithms.bounds() which is too slow 483 return np.array(self.dataset.GetBounds()) 484 485 def threshold(self, name=None, above=None, below=None, on="cells") -> Self: 486 """ 487 Threshold the tetrahedral mesh by a cell scalar value. 488 Reduce to only cells which satisfy the threshold limits. 489 490 - if `above = below` will only select cells with that specific value. 491 - if `above > below` selection range is flipped. 492 493 Set keyword "on" to either "cells" or "points". 494 """ 495 th = vtki.new("Threshold") 496 th.SetInputData(self.dataset) 497 498 if name is None: 499 if self.celldata.keys(): 500 name = self.celldata.keys()[0] 501 th.SetInputArrayToProcess(0, 0, 0, 1, name) 502 elif self.pointdata.keys(): 503 name = self.pointdata.keys()[0] 504 th.SetInputArrayToProcess(0, 0, 0, 0, name) 505 if name is None: 506 vedo.logger.warning("cannot find active array. Skip.") 507 return self 508 else: 509 if on.startswith("c"): 510 th.SetInputArrayToProcess(0, 0, 0, 1, name) 511 else: 512 th.SetInputArrayToProcess(0, 0, 0, 0, name) 513 514 if above is not None: 515 th.SetLowerThreshold(above) 516 517 if below is not None: 518 th.SetUpperThreshold(below) 519 520 th.Update() 521 return self._update(th.GetOutput()) 522 523 def isosurface(self, value=None, flying_edges=False) -> "vedo.mesh.Mesh": 524 """ 525 Return an `Mesh` isosurface extracted from the `Volume` object. 526 527 Set `value` as single float or list of values to draw the isosurface(s). 528 Use flying_edges for faster results (but sometimes can interfere with `smooth()`). 529 530 Examples: 531 - [isosurfaces1.py](https://github.com/marcomusy/vedo/tree/master/examples/volumetric/isosurfaces1.py) 532 533  534 """ 535 scrange = self.dataset.GetScalarRange() 536 537 if flying_edges: 538 cf = vtki.new("FlyingEdges3D") 539 cf.InterpolateAttributesOn() 540 else: 541 cf = vtki.new("ContourFilter") 542 cf.UseScalarTreeOn() 543 544 cf.SetInputData(self.dataset) 545 cf.ComputeNormalsOn() 546 547 if utils.is_sequence(value): 548 cf.SetNumberOfContours(len(value)) 549 for i, t in enumerate(value): 550 cf.SetValue(i, t) 551 else: 552 if value is None: 553 value = (2 * scrange[0] + scrange[1]) / 3.0 554 # print("automatic isosurface value =", value) 555 cf.SetValue(0, value) 556 557 cf.Update() 558 poly = cf.GetOutput() 559 560 out = vedo.mesh.Mesh(poly, c=None).flat() 561 out.mapper.SetScalarRange(scrange[0], scrange[1]) 562 563 out.pipeline = utils.OperationNode( 564 "isosurface", 565 parents=[self], 566 comment=f"#pts {out.dataset.GetNumberOfPoints()}", 567 c="#4cc9f0:#e9c46a", 568 ) 569 return out 570 571 def shrink(self, fraction=0.8) -> Self: 572 """ 573 Shrink the individual cells. 574 575  576 """ 577 sf = vtki.new("ShrinkFilter") 578 sf.SetInputData(self.dataset) 579 sf.SetShrinkFactor(fraction) 580 sf.Update() 581 out = sf.GetOutput() 582 self._update(out) 583 self.pipeline = utils.OperationNode( 584 "shrink", comment=f"by {fraction}", parents=[self], c="#9e2a2b" 585 ) 586 return self 587 588 def tomesh(self, fill=False, shrink=1.0) -> "vedo.mesh.Mesh": 589 """ 590 Build a polygonal `Mesh` from the current object. 591 592 If `fill=True`, the interior faces of all the cells are created. 593 (setting a `shrink` value slightly smaller than the default 1.0 594 can avoid flickering due to internal adjacent faces). 595 596 If `fill=False`, only the boundary faces will be generated. 597 """ 598 gf = vtki.new("GeometryFilter") 599 if fill: 600 sf = vtki.new("ShrinkFilter") 601 sf.SetInputData(self.dataset) 602 sf.SetShrinkFactor(shrink) 603 sf.Update() 604 gf.SetInputData(sf.GetOutput()) 605 gf.Update() 606 poly = gf.GetOutput() 607 else: 608 gf.SetInputData(self.dataset) 609 gf.Update() 610 poly = gf.GetOutput() 611 612 msh = vedo.mesh.Mesh(poly) 613 msh.copy_properties_from(self) 614 msh.pipeline = utils.OperationNode( 615 "tomesh", parents=[self], comment=f"fill={fill}", c="#9e2a2b:#e9c46a" 616 ) 617 return msh 618 619 @property 620 def cell_types_array(self): 621 """Return the list of cell types in the dataset.""" 622 uarr = self.dataset.GetCellTypesArray() 623 return utils.vtk2numpy(uarr) 624 625 def extract_cells_by_type(self, ctype) -> "UnstructuredGrid": 626 """Extract a specific cell type and return a new `UnstructuredGrid`.""" 627 if isinstance(ctype, str): 628 try: 629 ctype = vtki.cell_types[ctype.upper()] 630 except KeyError: 631 vedo.logger.error(f"extract_cells_by_type: cell type {ctype} does not exist. Skip.") 632 return self 633 uarr = self.dataset.GetCellTypesArray() 634 ctarrtyp = np.where(utils.vtk2numpy(uarr) == ctype)[0] 635 uarrtyp = utils.numpy2vtk(ctarrtyp, deep=False, dtype="id") 636 selection_node = vtki.new("SelectionNode") 637 selection_node.SetFieldType(vtki.get_class("SelectionNode").CELL) 638 selection_node.SetContentType(vtki.get_class("SelectionNode").INDICES) 639 selection_node.SetSelectionList(uarrtyp) 640 selection = vtki.new("Selection") 641 selection.AddNode(selection_node) 642 es = vtki.new("ExtractSelection") 643 es.SetInputData(0, self.dataset) 644 es.SetInputData(1, selection) 645 es.Update() 646 647 ug = UnstructuredGrid(es.GetOutput()) 648 ug.pipeline = utils.OperationNode( 649 "extract_cell_type", comment=f"type {ctype}", c="#edabab", parents=[self] 650 ) 651 return ug 652 653 def extract_cells_by_id(self, idlist, use_point_ids=False) -> "UnstructuredGrid": 654 """Return a new `UnstructuredGrid` composed of the specified subset of indices.""" 655 selection_node = vtki.new("SelectionNode") 656 if use_point_ids: 657 selection_node.SetFieldType(vtki.get_class("SelectionNode").POINT) 658 contcells = vtki.get_class("SelectionNode").CONTAINING_CELLS() 659 selection_node.GetProperties().Set(contcells, 1) 660 else: 661 selection_node.SetFieldType(vtki.get_class("SelectionNode").CELL) 662 selection_node.SetContentType(vtki.get_class("SelectionNode").INDICES) 663 vidlist = utils.numpy2vtk(idlist, dtype="id") 664 selection_node.SetSelectionList(vidlist) 665 selection = vtki.new("Selection") 666 selection.AddNode(selection_node) 667 es = vtki.new("ExtractSelection") 668 es.SetInputData(0, self) 669 es.SetInputData(1, selection) 670 es.Update() 671 672 ug = UnstructuredGrid(es.GetOutput()) 673 pr = vtki.vtkProperty() 674 pr.DeepCopy(self.properties) 675 ug.actor.SetProperty(pr) 676 ug.properties = pr 677 678 ug.mapper.SetLookupTable(utils.ctf2lut(self)) 679 ug.pipeline = utils.OperationNode( 680 "extract_cells_by_id", 681 parents=[self], 682 comment=f"#cells {self.dataset.GetNumberOfCells()}", 683 c="#9e2a2b", 684 ) 685 return ug 686 687 def find_cell(self, p: list) -> int: 688 """Locate the cell that contains a point and return the cell ID.""" 689 if self.cell_locator is None: 690 self.cell_locator = vtki.new("CellLocator") 691 self.cell_locator.SetDataSet(self.dataset) 692 self.cell_locator.BuildLocator() 693 cid = self.cell_locator.FindCell(p) 694 return cid 695 696 def clean(self) -> Self: 697 """ 698 Cleanup unused points and empty cells 699 """ 700 cl = vtki.new("StaticCleanUnstructuredGrid") 701 cl.SetInputData(self.dataset) 702 cl.RemoveUnusedPointsOn() 703 cl.ProduceMergeMapOff() 704 cl.AveragePointDataOff() 705 cl.Update() 706 707 self._update(cl.GetOutput()) 708 self.pipeline = utils.OperationNode( 709 "clean", 710 parents=[self], 711 comment=f"#cells {self.dataset.GetNumberOfCells()}", 712 c="#9e2a2b", 713 ) 714 return self 715 716 def extract_cells_on_plane(self, origin: tuple, normal: tuple) -> Self: 717 """ 718 Extract cells that are lying of the specified surface. 719 """ 720 bf = vtki.new("3DLinearGridCrinkleExtractor") 721 bf.SetInputData(self.dataset) 722 bf.CopyPointDataOn() 723 bf.CopyCellDataOn() 724 bf.RemoveUnusedPointsOff() 725 726 plane = vtki.new("Plane") 727 plane.SetOrigin(origin) 728 plane.SetNormal(normal) 729 bf.SetImplicitFunction(plane) 730 bf.Update() 731 732 self._update(bf.GetOutput(), reset_locators=False) 733 self.pipeline = utils.OperationNode( 734 "extract_cells_on_plane", 735 parents=[self], 736 comment=f"#cells {self.dataset.GetNumberOfCells()}", 737 c="#9e2a2b", 738 ) 739 return self 740 741 def extract_cells_on_sphere(self, center: tuple, radius: tuple) -> Self: 742 """ 743 Extract cells that are lying of the specified surface. 744 """ 745 bf = vtki.new("3DLinearGridCrinkleExtractor") 746 bf.SetInputData(self.dataset) 747 bf.CopyPointDataOn() 748 bf.CopyCellDataOn() 749 bf.RemoveUnusedPointsOff() 750 751 sph = vtki.new("Sphere") 752 sph.SetRadius(radius) 753 sph.SetCenter(center) 754 bf.SetImplicitFunction(sph) 755 bf.Update() 756 757 self._update(bf.GetOutput()) 758 self.pipeline = utils.OperationNode( 759 "extract_cells_on_sphere", 760 parents=[self], 761 comment=f"#cells {self.dataset.GetNumberOfCells()}", 762 c="#9e2a2b", 763 ) 764 return self 765 766 def extract_cells_on_cylinder(self, center: tuple, axis: tuple, radius: float) -> Self: 767 """ 768 Extract cells that are lying of the specified surface. 769 """ 770 bf = vtki.new("3DLinearGridCrinkleExtractor") 771 bf.SetInputData(self.dataset) 772 bf.CopyPointDataOn() 773 bf.CopyCellDataOn() 774 bf.RemoveUnusedPointsOff() 775 776 cyl = vtki.new("Cylinder") 777 cyl.SetRadius(radius) 778 cyl.SetCenter(center) 779 cyl.SetAxis(axis) 780 bf.SetImplicitFunction(cyl) 781 bf.Update() 782 783 self.pipeline = utils.OperationNode( 784 "extract_cells_on_cylinder", 785 parents=[self], 786 comment=f"#cells {self.dataset.GetNumberOfCells()}", 787 c="#9e2a2b", 788 ) 789 self._update(bf.GetOutput()) 790 return self 791 792 def cut_with_plane(self, origin=(0, 0, 0), normal="x") -> "UnstructuredGrid": 793 """ 794 Cut the object with the plane defined by a point and a normal. 795 796 Arguments: 797 origin : (list) 798 the cutting plane goes through this point 799 normal : (list, str) 800 normal vector to the cutting plane 801 """ 802 # if isinstance(self, vedo.Volume): 803 # raise RuntimeError("cut_with_plane() is not applicable to Volume objects.") 804 805 strn = str(normal) 806 if strn == "x": normal = (1, 0, 0) 807 elif strn == "y": normal = (0, 1, 0) 808 elif strn == "z": normal = (0, 0, 1) 809 elif strn == "-x": normal = (-1, 0, 0) 810 elif strn == "-y": normal = (0, -1, 0) 811 elif strn == "-z": normal = (0, 0, -1) 812 plane = vtki.new("Plane") 813 plane.SetOrigin(origin) 814 plane.SetNormal(normal) 815 clipper = vtki.new("ClipDataSet") 816 clipper.SetInputData(self.dataset) 817 clipper.SetClipFunction(plane) 818 clipper.GenerateClipScalarsOff() 819 clipper.GenerateClippedOutputOff() 820 clipper.SetValue(0) 821 clipper.Update() 822 cout = clipper.GetOutput() 823 824 if isinstance(cout, vtki.vtkUnstructuredGrid): 825 ug = vedo.UnstructuredGrid(cout) 826 if isinstance(self, vedo.UnstructuredGrid): 827 self._update(cout) 828 self.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 829 return self 830 ug.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 831 return ug 832 833 else: 834 self._update(cout) 835 self.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 836 return self 837 838 def cut_with_box(self, box: Any) -> "UnstructuredGrid": 839 """ 840 Cut the grid with the specified bounding box. 841 842 Parameter box has format [xmin, xmax, ymin, ymax, zmin, zmax]. 843 If an object is passed, its bounding box are used. 844 845 This method always returns a TetMesh object. 846 847 Example: 848 ```python 849 from vedo import * 850 tmesh = TetMesh(dataurl+'limb_ugrid.vtk') 851 tmesh.color('rainbow') 852 cu = Cube(side=500).x(500) # any Mesh works 853 tmesh.cut_with_box(cu).show(axes=1) 854 ``` 855 856  857 """ 858 bc = vtki.new("BoxClipDataSet") 859 bc.SetInputData(self.dataset) 860 try: 861 boxb = box.bounds() 862 except AttributeError: 863 boxb = box 864 865 bc.SetBoxClip(*boxb) 866 bc.Update() 867 cout = bc.GetOutput() 868 869 # output of vtkBoxClipDataSet is always tetrahedrons 870 tm = vedo.TetMesh(cout) 871 tm.pipeline = utils.OperationNode("cut_with_box", parents=[self], c="#9e2a2b") 872 return tm 873 874 def cut_with_mesh(self, mesh: Mesh, invert=False, whole_cells=False, on_boundary=False) -> "UnstructuredGrid": 875 """ 876 Cut a `UnstructuredGrid` or `TetMesh` with a `Mesh`. 877 878 Use `invert` to return cut off part of the input object. 879 """ 880 ug = self.dataset 881 882 ippd = vtki.new("ImplicitPolyDataDistance") 883 ippd.SetInput(mesh.dataset) 884 885 if whole_cells or on_boundary: 886 clipper = vtki.new("ExtractGeometry") 887 clipper.SetInputData(ug) 888 clipper.SetImplicitFunction(ippd) 889 clipper.SetExtractInside(not invert) 890 clipper.SetExtractBoundaryCells(False) 891 if on_boundary: 892 clipper.SetExtractBoundaryCells(True) 893 clipper.SetExtractOnlyBoundaryCells(True) 894 else: 895 signed_dists = vtki.vtkFloatArray() 896 signed_dists.SetNumberOfComponents(1) 897 signed_dists.SetName("SignedDistance") 898 for pointId in range(ug.GetNumberOfPoints()): 899 p = ug.GetPoint(pointId) 900 signed_dist = ippd.EvaluateFunction(p) 901 signed_dists.InsertNextValue(signed_dist) 902 ug.GetPointData().AddArray(signed_dists) 903 ug.GetPointData().SetActiveScalars("SignedDistance") # NEEDED 904 clipper = vtki.new("ClipDataSet") 905 clipper.SetInputData(ug) 906 clipper.SetInsideOut(not invert) 907 clipper.SetValue(0.0) 908 909 clipper.Update() 910 911 out = vedo.UnstructuredGrid(clipper.GetOutput()) 912 out.pipeline = utils.OperationNode("cut_with_mesh", parents=[self], c="#9e2a2b") 913 return out
Support for UnstructuredGrid objects.
41 def __init__(self, inputobj=None): 42 """ 43 Support for UnstructuredGrid objects. 44 45 Arguments: 46 inputobj : (list, vtkUnstructuredGrid, str) 47 A list in the form `[points, cells, celltypes]`, 48 or a vtkUnstructuredGrid object, or a filename 49 50 Celltypes are identified by the following 51 [convention](https://vtk.org/doc/nightly/html/vtkCellType_8h_source.html). 52 """ 53 super().__init__() 54 55 self.dataset = None 56 57 self.mapper = vtki.new("PolyDataMapper") 58 self._actor = vtki.vtkActor() 59 self._actor.retrieve_object = weak_ref_to(self) 60 self._actor.SetMapper(self.mapper) 61 self.properties = self._actor.GetProperty() 62 63 self.transform = LinearTransform() 64 self.point_locator = None 65 self.cell_locator = None 66 self.line_locator = None 67 68 self.name = "UnstructuredGrid" 69 self.filename = "" 70 self.file_size = "" 71 72 self.info = {} 73 self.time = time.time() 74 self.rendered_at = set() 75 76 ###################################### 77 inputtype = str(type(inputobj)) 78 79 if inputobj is None: 80 self.dataset = vtki.vtkUnstructuredGrid() 81 82 elif utils.is_sequence(inputobj): 83 84 pts, cells, celltypes = inputobj 85 assert len(cells) == len(celltypes) 86 87 self.dataset = vtki.vtkUnstructuredGrid() 88 89 if not utils.is_sequence(cells[0]): 90 tets = [] 91 nf = cells[0] + 1 92 for i, cl in enumerate(cells): 93 if i in (nf, 0): 94 k = i + 1 95 nf = cl + k 96 cell = [cells[j + k] for j in range(cl)] 97 tets.append(cell) 98 cells = tets 99 100 # This would fill the points and use those to define orientation 101 vpts = utils.numpy2vtk(pts, dtype=np.float32) 102 points = vtki.vtkPoints() 103 points.SetData(vpts) 104 self.dataset.SetPoints(points) 105 106 # Fill cells 107 # https://vtk.org/doc/nightly/html/vtkCellType_8h_source.html 108 for i, ct in enumerate(celltypes): 109 if ct == vtki.cell_types["VERTEX"]: 110 cell = vtki.vtkVertex() 111 elif ct == vtki.cell_types["POLY_VERTEX"]: 112 cell = vtki.vtkPolyVertex() 113 elif ct == vtki.cell_types["TETRA"]: 114 cell = vtki.vtkTetra() 115 elif ct == vtki.cell_types["WEDGE"]: 116 cell = vtki.vtkWedge() 117 elif ct == vtki.cell_types["LINE"]: 118 cell = vtki.vtkLine() 119 elif ct == vtki.cell_types["POLY_LINE"]: 120 cell = vtki.vtkPolyLine() 121 elif ct == vtki.cell_types["TRIANGLE"]: 122 cell = vtki.vtkTriangle() 123 elif ct == vtki.cell_types["TRIANGLE_STRIP"]: 124 cell = vtki.vtkTriangleStrip() 125 elif ct == vtki.cell_types["POLYGON"]: 126 cell = vtki.vtkPolygon() 127 elif ct == vtki.cell_types["PIXEL"]: 128 cell = vtki.vtkPixel() 129 elif ct == vtki.cell_types["QUAD"]: 130 cell = vtki.vtkQuad() 131 elif ct == vtki.cell_types["VOXEL"]: 132 cell = vtki.vtkVoxel() 133 elif ct == vtki.cell_types["PYRAMID"]: 134 cell = vtki.vtkPyramid() 135 elif ct == vtki.cell_types["HEXAHEDRON"]: 136 cell = vtki.vtkHexahedron() 137 elif ct == vtki.cell_types["HEXAGONAL_PRISM"]: 138 cell = vtki.vtkHexagonalPrism() 139 elif ct == vtki.cell_types["PENTAGONAL_PRISM"]: 140 cell = vtki.vtkPentagonalPrism() 141 elif ct == vtki.cell_types["QUADRATIC_TETRA"]: 142 from vtkmodules.vtkCommonDataModel import vtkQuadraticTetra 143 cell = vtkQuadraticTetra() 144 elif ct == vtki.cell_types["QUADRATIC_HEXAHEDRON"]: 145 from vtkmodules.vtkCommonDataModel import vtkQuadraticHexahedron 146 cell = vtkQuadraticHexahedron() 147 elif ct == vtki.cell_types["QUADRATIC_WEDGE"]: 148 from vtkmodules.vtkCommonDataModel import vtkQuadraticWedge 149 cell = vtkQuadraticWedge() 150 elif ct == vtki.cell_types["QUADRATIC_PYRAMID"]: 151 from vtkmodules.vtkCommonDataModel import vtkQuadraticPyramid 152 cell = vtkQuadraticPyramid() 153 elif ct == vtki.cell_types["QUADRATIC_LINEAR_QUAD"]: 154 from vtkmodules.vtkCommonDataModel import vtkQuadraticLinearQuad 155 cell = vtkQuadraticLinearQuad() 156 elif ct == vtki.cell_types["QUADRATIC_LINEAR_WEDGE"]: 157 from vtkmodules.vtkCommonDataModel import vtkQuadraticLinearWedge 158 cell = vtkQuadraticLinearWedge() 159 elif ct == vtki.cell_types["BIQUADRATIC_QUADRATIC_WEDGE"]: 160 from vtkmodules.vtkCommonDataModel import vtkBiQuadraticQuadraticWedge 161 cell = vtkBiQuadraticQuadraticWedge() 162 elif ct == vtki.cell_types["BIQUADRATIC_QUADRATIC_HEXAHEDRON"]: 163 from vtkmodules.vtkCommonDataModel import vtkBiQuadraticQuadraticHexahedron 164 cell = vtkBiQuadraticQuadraticHexahedron() 165 elif ct == vtki.cell_types["BIQUADRATIC_TRIANGLE"]: 166 from vtkmodules.vtkCommonDataModel import vtkBiQuadraticTriangle 167 cell = vtkBiQuadraticTriangle() 168 elif ct == vtki.cell_types["CUBIC_LINE"]: 169 from vtkmodules.vtkCommonDataModel import vtkCubicLine 170 cell = vtkCubicLine() 171 elif ct == vtki.cell_types["CONVEX_POINT_SET"]: 172 from vtkmodules.vtkCommonDataModel import vtkConvexPointSet 173 cell = vtkConvexPointSet() 174 elif ct == vtki.cell_types["POLYHEDRON"]: 175 from vtkmodules.vtkCommonDataModel import vtkPolyhedron 176 cell = vtkPolyhedron() 177 elif ct == vtki.cell_types["HIGHER_ORDER_TRIANGLE"]: 178 from vtkmodules.vtkCommonDataModel import vtkHigherOrderTriangle 179 cell = vtkHigherOrderTriangle() 180 elif ct == vtki.cell_types["HIGHER_ORDER_QUAD"]: 181 from vtkmodules.vtkCommonDataModel import vtkHigherOrderQuadrilateral 182 cell = vtkHigherOrderQuadrilateral() 183 elif ct == vtki.cell_types["HIGHER_ORDER_TETRAHEDRON"]: 184 from vtkmodules.vtkCommonDataModel import vtkHigherOrderTetra 185 cell = vtkHigherOrderTetra() 186 elif ct == vtki.cell_types["HIGHER_ORDER_WEDGE"]: 187 from vtkmodules.vtkCommonDataModel import vtkHigherOrderWedge 188 cell = vtkHigherOrderWedge() 189 elif ct == vtki.cell_types["HIGHER_ORDER_HEXAHEDRON"]: 190 from vtkmodules.vtkCommonDataModel import vtkHigherOrderHexahedron 191 cell = vtkHigherOrderHexahedron() 192 elif ct == vtki.cell_types["LAGRANGE_CURVE"]: 193 from vtkmodules.vtkCommonDataModel import vtkLagrangeCurve 194 cell = vtkLagrangeCurve() 195 elif ct == vtki.cell_types["LAGRANGE_TRIANGLE"]: 196 from vtkmodules.vtkCommonDataModel import vtkLagrangeTriangle 197 cell = vtkLagrangeTriangle() 198 elif ct == vtki.cell_types["LAGRANGE_QUADRILATERAL"]: 199 from vtkmodules.vtkCommonDataModel import vtkLagrangeQuadrilateral 200 cell = vtkLagrangeQuadrilateral() 201 elif ct == vtki.cell_types["LAGRANGE_TETRAHEDRON"]: 202 from vtkmodules.vtkCommonDataModel import vtkLagrangeTetra 203 cell = vtkLagrangeTetra() 204 elif ct == vtki.cell_types["LAGRANGE_HEXAHEDRON"]: 205 from vtkmodules.vtkCommonDataModel import vtkLagrangeHexahedron 206 cell = vtkLagrangeHexahedron() 207 elif ct == vtki.cell_types["LAGRANGE_WEDGE"]: 208 from vtkmodules.vtkCommonDataModel import vtkLagrangeWedge 209 cell = vtkLagrangeWedge() 210 elif ct == vtki.cell_types["BEZIER_CURVE"]: 211 from vtkmodules.vtkCommonDataModel import vtkBezierCurve 212 cell = vtkBezierCurve() 213 elif ct == vtki.cell_types["BEZIER_TRIANGLE"]: 214 from vtkmodules.vtkCommonDataModel import vtkBezierTriangle 215 cell = vtkBezierTriangle() 216 elif ct == vtki.cell_types["BEZIER_QUADRILATERAL"]: 217 from vtkmodules.vtkCommonDataModel import vtkBezierQuadrilateral 218 cell = vtkBezierQuadrilateral() 219 elif ct == vtki.cell_types["BEZIER_TETRAHEDRON"]: 220 from vtkmodules.vtkCommonDataModel import vtkBezierTetra 221 cell = vtkBezierTetra() 222 elif ct == vtki.cell_types["BEZIER_HEXAHEDRON"]: 223 from vtkmodules.vtkCommonDataModel import vtkBezierHexahedron 224 cell = vtkBezierHexahedron() 225 elif ct == vtki.cell_types["BEZIER_WEDGE"]: 226 from vtkmodules.vtkCommonDataModel import vtkBezierWedge 227 cell = vtkBezierWedge() 228 else: 229 vedo.logger.error( 230 f"UnstructuredGrid: cell type {ct} not supported. Skip.") 231 continue 232 233 cpids = cell.GetPointIds() 234 cell_conn = cells[i] 235 for j, pid in enumerate(cell_conn): 236 cpids.SetId(j, pid) 237 self.dataset.InsertNextCell(ct, cpids) 238 239 elif "UnstructuredGrid" in inputtype: 240 self.dataset = inputobj 241 242 elif isinstance(inputobj, str): 243 if "https://" in inputobj: 244 inputobj = download(inputobj, verbose=False) 245 self.filename = inputobj 246 if inputobj.endswith(".vtu"): 247 reader = vtki.new("XMLUnstructuredGridReader") 248 else: 249 reader = vtki.new("UnstructuredGridReader") 250 self.filename = inputobj 251 reader.SetFileName(inputobj) 252 reader.Update() 253 self.dataset = reader.GetOutput() 254 255 else: 256 # this converts other types of vtk objects to UnstructuredGrid 257 apf = vtki.new("AppendFilter") 258 try: 259 apf.AddInputData(inputobj) 260 except TypeError: 261 apf.AddInputData(inputobj.dataset) 262 apf.Update() 263 self.dataset = apf.GetOutput() 264 265 self.properties.SetColor(0.89, 0.455, 0.671) # pink7 266 267 self.pipeline = utils.OperationNode( 268 self, comment=f"#cells {self.dataset.GetNumberOfCells()}", c="#4cc9f0" 269 )
Support for UnstructuredGrid objects.
Arguments:
- inputobj : (list, vtkUnstructuredGrid, str)
A list in the form
[points, cells, celltypes], or a vtkUnstructuredGrid object, or a filename
Celltypes are identified by the following convention.
412 @property 413 def actor(self): 414 """Return the `vtkActor` of the object.""" 415 # print("building actor") 416 gf = vtki.new("GeometryFilter") 417 gf.SetInputData(self.dataset) 418 gf.Update() 419 out = gf.GetOutput() 420 self.mapper.SetInputData(out) 421 self.mapper.Modified() 422 return self._actor
Return the vtkActor of the object.
435 def merge(self, *others) -> Self: 436 """ 437 Merge multiple datasets into one single `UnstrcturedGrid`. 438 """ 439 apf = vtki.new("AppendFilter") 440 for o in others: 441 if isinstance(o, UnstructuredGrid): 442 apf.AddInputData(o.dataset) 443 elif isinstance(o, vtki.vtkUnstructuredGrid): 444 apf.AddInputData(o) 445 else: 446 vedo.printc("Error: cannot merge type", type(o), c="r") 447 apf.Update() 448 self._update(apf.GetOutput()) 449 self.pipeline = utils.OperationNode( 450 "merge", parents=[self, *others], c="#9e2a2b" 451 ) 452 return self
Merge multiple datasets into one single UnstrcturedGrid.
454 def copy(self, deep=True) -> "UnstructuredGrid": 455 """Return a copy of the object. Alias of `clone()`.""" 456 return self.clone(deep=deep)
Return a copy of the object. Alias of clone().
458 def clone(self, deep=True) -> "UnstructuredGrid": 459 """Clone the UnstructuredGrid object to yield an exact copy.""" 460 ug = vtki.vtkUnstructuredGrid() 461 if deep: 462 ug.DeepCopy(self.dataset) 463 else: 464 ug.ShallowCopy(self.dataset) 465 if isinstance(self, vedo.UnstructuredGrid): 466 cloned = vedo.UnstructuredGrid(ug) 467 else: 468 cloned = vedo.TetMesh(ug) 469 470 cloned.copy_properties_from(self) 471 472 cloned.pipeline = utils.OperationNode( 473 "clone", parents=[self], shape="diamond", c="#bbe1ed" 474 ) 475 return cloned
Clone the UnstructuredGrid object to yield an exact copy.
477 def bounds(self) -> np.ndarray: 478 """ 479 Get the object bounds. 480 Returns a list in format `[xmin,xmax, ymin,ymax, zmin,zmax]`. 481 """ 482 # OVERRIDE CommonAlgorithms.bounds() which is too slow 483 return np.array(self.dataset.GetBounds())
Get the object bounds.
Returns a list in format [xmin,xmax, ymin,ymax, zmin,zmax].
485 def threshold(self, name=None, above=None, below=None, on="cells") -> Self: 486 """ 487 Threshold the tetrahedral mesh by a cell scalar value. 488 Reduce to only cells which satisfy the threshold limits. 489 490 - if `above = below` will only select cells with that specific value. 491 - if `above > below` selection range is flipped. 492 493 Set keyword "on" to either "cells" or "points". 494 """ 495 th = vtki.new("Threshold") 496 th.SetInputData(self.dataset) 497 498 if name is None: 499 if self.celldata.keys(): 500 name = self.celldata.keys()[0] 501 th.SetInputArrayToProcess(0, 0, 0, 1, name) 502 elif self.pointdata.keys(): 503 name = self.pointdata.keys()[0] 504 th.SetInputArrayToProcess(0, 0, 0, 0, name) 505 if name is None: 506 vedo.logger.warning("cannot find active array. Skip.") 507 return self 508 else: 509 if on.startswith("c"): 510 th.SetInputArrayToProcess(0, 0, 0, 1, name) 511 else: 512 th.SetInputArrayToProcess(0, 0, 0, 0, name) 513 514 if above is not None: 515 th.SetLowerThreshold(above) 516 517 if below is not None: 518 th.SetUpperThreshold(below) 519 520 th.Update() 521 return self._update(th.GetOutput())
Threshold the tetrahedral mesh by a cell scalar value. Reduce to only cells which satisfy the threshold limits.
- if
above = belowwill only select cells with that specific value. - if
above > belowselection range is flipped.
Set keyword "on" to either "cells" or "points".
523 def isosurface(self, value=None, flying_edges=False) -> "vedo.mesh.Mesh": 524 """ 525 Return an `Mesh` isosurface extracted from the `Volume` object. 526 527 Set `value` as single float or list of values to draw the isosurface(s). 528 Use flying_edges for faster results (but sometimes can interfere with `smooth()`). 529 530 Examples: 531 - [isosurfaces1.py](https://github.com/marcomusy/vedo/tree/master/examples/volumetric/isosurfaces1.py) 532 533  534 """ 535 scrange = self.dataset.GetScalarRange() 536 537 if flying_edges: 538 cf = vtki.new("FlyingEdges3D") 539 cf.InterpolateAttributesOn() 540 else: 541 cf = vtki.new("ContourFilter") 542 cf.UseScalarTreeOn() 543 544 cf.SetInputData(self.dataset) 545 cf.ComputeNormalsOn() 546 547 if utils.is_sequence(value): 548 cf.SetNumberOfContours(len(value)) 549 for i, t in enumerate(value): 550 cf.SetValue(i, t) 551 else: 552 if value is None: 553 value = (2 * scrange[0] + scrange[1]) / 3.0 554 # print("automatic isosurface value =", value) 555 cf.SetValue(0, value) 556 557 cf.Update() 558 poly = cf.GetOutput() 559 560 out = vedo.mesh.Mesh(poly, c=None).flat() 561 out.mapper.SetScalarRange(scrange[0], scrange[1]) 562 563 out.pipeline = utils.OperationNode( 564 "isosurface", 565 parents=[self], 566 comment=f"#pts {out.dataset.GetNumberOfPoints()}", 567 c="#4cc9f0:#e9c46a", 568 ) 569 return out
Return an Mesh isosurface extracted from the Volume object.
Set value as single float or list of values to draw the isosurface(s).
Use flying_edges for faster results (but sometimes can interfere with smooth()).
Examples:
571 def shrink(self, fraction=0.8) -> Self: 572 """ 573 Shrink the individual cells. 574 575  576 """ 577 sf = vtki.new("ShrinkFilter") 578 sf.SetInputData(self.dataset) 579 sf.SetShrinkFactor(fraction) 580 sf.Update() 581 out = sf.GetOutput() 582 self._update(out) 583 self.pipeline = utils.OperationNode( 584 "shrink", comment=f"by {fraction}", parents=[self], c="#9e2a2b" 585 ) 586 return self
Shrink the individual cells.

588 def tomesh(self, fill=False, shrink=1.0) -> "vedo.mesh.Mesh": 589 """ 590 Build a polygonal `Mesh` from the current object. 591 592 If `fill=True`, the interior faces of all the cells are created. 593 (setting a `shrink` value slightly smaller than the default 1.0 594 can avoid flickering due to internal adjacent faces). 595 596 If `fill=False`, only the boundary faces will be generated. 597 """ 598 gf = vtki.new("GeometryFilter") 599 if fill: 600 sf = vtki.new("ShrinkFilter") 601 sf.SetInputData(self.dataset) 602 sf.SetShrinkFactor(shrink) 603 sf.Update() 604 gf.SetInputData(sf.GetOutput()) 605 gf.Update() 606 poly = gf.GetOutput() 607 else: 608 gf.SetInputData(self.dataset) 609 gf.Update() 610 poly = gf.GetOutput() 611 612 msh = vedo.mesh.Mesh(poly) 613 msh.copy_properties_from(self) 614 msh.pipeline = utils.OperationNode( 615 "tomesh", parents=[self], comment=f"fill={fill}", c="#9e2a2b:#e9c46a" 616 ) 617 return msh
Build a polygonal Mesh from the current object.
If fill=True, the interior faces of all the cells are created.
(setting a shrink value slightly smaller than the default 1.0
can avoid flickering due to internal adjacent faces).
If fill=False, only the boundary faces will be generated.
619 @property 620 def cell_types_array(self): 621 """Return the list of cell types in the dataset.""" 622 uarr = self.dataset.GetCellTypesArray() 623 return utils.vtk2numpy(uarr)
Return the list of cell types in the dataset.
625 def extract_cells_by_type(self, ctype) -> "UnstructuredGrid": 626 """Extract a specific cell type and return a new `UnstructuredGrid`.""" 627 if isinstance(ctype, str): 628 try: 629 ctype = vtki.cell_types[ctype.upper()] 630 except KeyError: 631 vedo.logger.error(f"extract_cells_by_type: cell type {ctype} does not exist. Skip.") 632 return self 633 uarr = self.dataset.GetCellTypesArray() 634 ctarrtyp = np.where(utils.vtk2numpy(uarr) == ctype)[0] 635 uarrtyp = utils.numpy2vtk(ctarrtyp, deep=False, dtype="id") 636 selection_node = vtki.new("SelectionNode") 637 selection_node.SetFieldType(vtki.get_class("SelectionNode").CELL) 638 selection_node.SetContentType(vtki.get_class("SelectionNode").INDICES) 639 selection_node.SetSelectionList(uarrtyp) 640 selection = vtki.new("Selection") 641 selection.AddNode(selection_node) 642 es = vtki.new("ExtractSelection") 643 es.SetInputData(0, self.dataset) 644 es.SetInputData(1, selection) 645 es.Update() 646 647 ug = UnstructuredGrid(es.GetOutput()) 648 ug.pipeline = utils.OperationNode( 649 "extract_cell_type", comment=f"type {ctype}", c="#edabab", parents=[self] 650 ) 651 return ug
Extract a specific cell type and return a new UnstructuredGrid.
653 def extract_cells_by_id(self, idlist, use_point_ids=False) -> "UnstructuredGrid": 654 """Return a new `UnstructuredGrid` composed of the specified subset of indices.""" 655 selection_node = vtki.new("SelectionNode") 656 if use_point_ids: 657 selection_node.SetFieldType(vtki.get_class("SelectionNode").POINT) 658 contcells = vtki.get_class("SelectionNode").CONTAINING_CELLS() 659 selection_node.GetProperties().Set(contcells, 1) 660 else: 661 selection_node.SetFieldType(vtki.get_class("SelectionNode").CELL) 662 selection_node.SetContentType(vtki.get_class("SelectionNode").INDICES) 663 vidlist = utils.numpy2vtk(idlist, dtype="id") 664 selection_node.SetSelectionList(vidlist) 665 selection = vtki.new("Selection") 666 selection.AddNode(selection_node) 667 es = vtki.new("ExtractSelection") 668 es.SetInputData(0, self) 669 es.SetInputData(1, selection) 670 es.Update() 671 672 ug = UnstructuredGrid(es.GetOutput()) 673 pr = vtki.vtkProperty() 674 pr.DeepCopy(self.properties) 675 ug.actor.SetProperty(pr) 676 ug.properties = pr 677 678 ug.mapper.SetLookupTable(utils.ctf2lut(self)) 679 ug.pipeline = utils.OperationNode( 680 "extract_cells_by_id", 681 parents=[self], 682 comment=f"#cells {self.dataset.GetNumberOfCells()}", 683 c="#9e2a2b", 684 ) 685 return ug
Return a new UnstructuredGrid composed of the specified subset of indices.
687 def find_cell(self, p: list) -> int: 688 """Locate the cell that contains a point and return the cell ID.""" 689 if self.cell_locator is None: 690 self.cell_locator = vtki.new("CellLocator") 691 self.cell_locator.SetDataSet(self.dataset) 692 self.cell_locator.BuildLocator() 693 cid = self.cell_locator.FindCell(p) 694 return cid
Locate the cell that contains a point and return the cell ID.
696 def clean(self) -> Self: 697 """ 698 Cleanup unused points and empty cells 699 """ 700 cl = vtki.new("StaticCleanUnstructuredGrid") 701 cl.SetInputData(self.dataset) 702 cl.RemoveUnusedPointsOn() 703 cl.ProduceMergeMapOff() 704 cl.AveragePointDataOff() 705 cl.Update() 706 707 self._update(cl.GetOutput()) 708 self.pipeline = utils.OperationNode( 709 "clean", 710 parents=[self], 711 comment=f"#cells {self.dataset.GetNumberOfCells()}", 712 c="#9e2a2b", 713 ) 714 return self
Cleanup unused points and empty cells
716 def extract_cells_on_plane(self, origin: tuple, normal: tuple) -> Self: 717 """ 718 Extract cells that are lying of the specified surface. 719 """ 720 bf = vtki.new("3DLinearGridCrinkleExtractor") 721 bf.SetInputData(self.dataset) 722 bf.CopyPointDataOn() 723 bf.CopyCellDataOn() 724 bf.RemoveUnusedPointsOff() 725 726 plane = vtki.new("Plane") 727 plane.SetOrigin(origin) 728 plane.SetNormal(normal) 729 bf.SetImplicitFunction(plane) 730 bf.Update() 731 732 self._update(bf.GetOutput(), reset_locators=False) 733 self.pipeline = utils.OperationNode( 734 "extract_cells_on_plane", 735 parents=[self], 736 comment=f"#cells {self.dataset.GetNumberOfCells()}", 737 c="#9e2a2b", 738 ) 739 return self
Extract cells that are lying of the specified surface.
741 def extract_cells_on_sphere(self, center: tuple, radius: tuple) -> Self: 742 """ 743 Extract cells that are lying of the specified surface. 744 """ 745 bf = vtki.new("3DLinearGridCrinkleExtractor") 746 bf.SetInputData(self.dataset) 747 bf.CopyPointDataOn() 748 bf.CopyCellDataOn() 749 bf.RemoveUnusedPointsOff() 750 751 sph = vtki.new("Sphere") 752 sph.SetRadius(radius) 753 sph.SetCenter(center) 754 bf.SetImplicitFunction(sph) 755 bf.Update() 756 757 self._update(bf.GetOutput()) 758 self.pipeline = utils.OperationNode( 759 "extract_cells_on_sphere", 760 parents=[self], 761 comment=f"#cells {self.dataset.GetNumberOfCells()}", 762 c="#9e2a2b", 763 ) 764 return self
Extract cells that are lying of the specified surface.
766 def extract_cells_on_cylinder(self, center: tuple, axis: tuple, radius: float) -> Self: 767 """ 768 Extract cells that are lying of the specified surface. 769 """ 770 bf = vtki.new("3DLinearGridCrinkleExtractor") 771 bf.SetInputData(self.dataset) 772 bf.CopyPointDataOn() 773 bf.CopyCellDataOn() 774 bf.RemoveUnusedPointsOff() 775 776 cyl = vtki.new("Cylinder") 777 cyl.SetRadius(radius) 778 cyl.SetCenter(center) 779 cyl.SetAxis(axis) 780 bf.SetImplicitFunction(cyl) 781 bf.Update() 782 783 self.pipeline = utils.OperationNode( 784 "extract_cells_on_cylinder", 785 parents=[self], 786 comment=f"#cells {self.dataset.GetNumberOfCells()}", 787 c="#9e2a2b", 788 ) 789 self._update(bf.GetOutput()) 790 return self
Extract cells that are lying of the specified surface.
792 def cut_with_plane(self, origin=(0, 0, 0), normal="x") -> "UnstructuredGrid": 793 """ 794 Cut the object with the plane defined by a point and a normal. 795 796 Arguments: 797 origin : (list) 798 the cutting plane goes through this point 799 normal : (list, str) 800 normal vector to the cutting plane 801 """ 802 # if isinstance(self, vedo.Volume): 803 # raise RuntimeError("cut_with_plane() is not applicable to Volume objects.") 804 805 strn = str(normal) 806 if strn == "x": normal = (1, 0, 0) 807 elif strn == "y": normal = (0, 1, 0) 808 elif strn == "z": normal = (0, 0, 1) 809 elif strn == "-x": normal = (-1, 0, 0) 810 elif strn == "-y": normal = (0, -1, 0) 811 elif strn == "-z": normal = (0, 0, -1) 812 plane = vtki.new("Plane") 813 plane.SetOrigin(origin) 814 plane.SetNormal(normal) 815 clipper = vtki.new("ClipDataSet") 816 clipper.SetInputData(self.dataset) 817 clipper.SetClipFunction(plane) 818 clipper.GenerateClipScalarsOff() 819 clipper.GenerateClippedOutputOff() 820 clipper.SetValue(0) 821 clipper.Update() 822 cout = clipper.GetOutput() 823 824 if isinstance(cout, vtki.vtkUnstructuredGrid): 825 ug = vedo.UnstructuredGrid(cout) 826 if isinstance(self, vedo.UnstructuredGrid): 827 self._update(cout) 828 self.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 829 return self 830 ug.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 831 return ug 832 833 else: 834 self._update(cout) 835 self.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 836 return self
Cut the object with the plane defined by a point and a normal.
Arguments:
- origin : (list) the cutting plane goes through this point
- normal : (list, str) normal vector to the cutting plane
838 def cut_with_box(self, box: Any) -> "UnstructuredGrid": 839 """ 840 Cut the grid with the specified bounding box. 841 842 Parameter box has format [xmin, xmax, ymin, ymax, zmin, zmax]. 843 If an object is passed, its bounding box are used. 844 845 This method always returns a TetMesh object. 846 847 Example: 848 ```python 849 from vedo import * 850 tmesh = TetMesh(dataurl+'limb_ugrid.vtk') 851 tmesh.color('rainbow') 852 cu = Cube(side=500).x(500) # any Mesh works 853 tmesh.cut_with_box(cu).show(axes=1) 854 ``` 855 856  857 """ 858 bc = vtki.new("BoxClipDataSet") 859 bc.SetInputData(self.dataset) 860 try: 861 boxb = box.bounds() 862 except AttributeError: 863 boxb = box 864 865 bc.SetBoxClip(*boxb) 866 bc.Update() 867 cout = bc.GetOutput() 868 869 # output of vtkBoxClipDataSet is always tetrahedrons 870 tm = vedo.TetMesh(cout) 871 tm.pipeline = utils.OperationNode("cut_with_box", parents=[self], c="#9e2a2b") 872 return tm
Cut the grid with the specified bounding box.
Parameter box has format [xmin, xmax, ymin, ymax, zmin, zmax]. If an object is passed, its bounding box are used.
This method always returns a TetMesh object.
Example:
from vedo import *
tmesh = TetMesh(dataurl+'limb_ugrid.vtk')
tmesh.color('rainbow')
cu = Cube(side=500).x(500) # any Mesh works
tmesh.cut_with_box(cu).show(axes=1)

874 def cut_with_mesh(self, mesh: Mesh, invert=False, whole_cells=False, on_boundary=False) -> "UnstructuredGrid": 875 """ 876 Cut a `UnstructuredGrid` or `TetMesh` with a `Mesh`. 877 878 Use `invert` to return cut off part of the input object. 879 """ 880 ug = self.dataset 881 882 ippd = vtki.new("ImplicitPolyDataDistance") 883 ippd.SetInput(mesh.dataset) 884 885 if whole_cells or on_boundary: 886 clipper = vtki.new("ExtractGeometry") 887 clipper.SetInputData(ug) 888 clipper.SetImplicitFunction(ippd) 889 clipper.SetExtractInside(not invert) 890 clipper.SetExtractBoundaryCells(False) 891 if on_boundary: 892 clipper.SetExtractBoundaryCells(True) 893 clipper.SetExtractOnlyBoundaryCells(True) 894 else: 895 signed_dists = vtki.vtkFloatArray() 896 signed_dists.SetNumberOfComponents(1) 897 signed_dists.SetName("SignedDistance") 898 for pointId in range(ug.GetNumberOfPoints()): 899 p = ug.GetPoint(pointId) 900 signed_dist = ippd.EvaluateFunction(p) 901 signed_dists.InsertNextValue(signed_dist) 902 ug.GetPointData().AddArray(signed_dists) 903 ug.GetPointData().SetActiveScalars("SignedDistance") # NEEDED 904 clipper = vtki.new("ClipDataSet") 905 clipper.SetInputData(ug) 906 clipper.SetInsideOut(not invert) 907 clipper.SetValue(0.0) 908 909 clipper.Update() 910 911 out = vedo.UnstructuredGrid(clipper.GetOutput()) 912 out.pipeline = utils.OperationNode("cut_with_mesh", parents=[self], c="#9e2a2b") 913 return out
Cut a UnstructuredGrid or TetMesh with a Mesh.
Use invert to return cut off part of the input object.
Inherited Members
- vedo.core.PointAlgorithms
- apply_transform
- apply_transform_from_actor
- pos
- shift
- x
- y
- z
- rotate
- rotate_x
- rotate_y
- rotate_z
- reorient
- scale
- vedo.core.CommonAlgorithms
- pointdata
- celldata
- metadata
- rename
- memory_address
- memory_size
- modified
- box
- update_dataset
- xbounds
- ybounds
- zbounds
- diagonal_size
- average_size
- center_of_mass
- copy_data_from
- inputdata
- npoints
- nvertices
- ncells
- cell_centers
- lines
- lines_as_flat_array
- mark_boundaries
- find_cells_in_bounds
- find_cells_along_line
- find_cells_along_plane
- keep_cell_types
- map_cells_to_points
- vertices
- points
- coordinates
- cells_as_flat_array
- cells
- cell_edge_neighbors
- map_points_to_cells
- resample_data_from
- interpolate_data_from
- add_ids
- gradient
- divergence
- vorticity
- probe
- compute_cell_size
- generate_random_data
- integrate_data
- write
- signed_distance
- unsigned_distance
- smooth_data
- compute_streamlines
- vedo.visual.MeshVisual
- follow_camera
- wireframe
- flat
- phong
- backface_culling
- render_lines_as_tubes
- frontface_culling
- backcolor
- bc
- linewidth
- lw
- linecolor
- lc
- texture
- vedo.visual.PointsVisual
- clone2d
- copy_properties_from
- color
- c
- alpha
- lut_color_at
- opacity
- force_opaque
- force_translucent
- point_size
- ps
- render_points_as_spheres
- lighting
- point_blurring
- cellcolors
- pointcolors
- cmap
- add_trail
- update_trail
- add_shadow
- update_shadows
- labels
- labels2d
- legend
- flagpole
- flagpost
- caption
917class TetMesh(UnstructuredGrid): 918 """The class describing tetrahedral meshes.""" 919 920 def __init__(self, inputobj=None): 921 """ 922 Arguments: 923 inputobj : (vtkUnstructuredGrid, list, str, tetgenpy.TetgenIO) 924 list of points and tet indices, or filename 925 """ 926 super().__init__() 927 928 self.dataset = None 929 930 self.mapper = vtki.new("PolyDataMapper") 931 self._actor = vtki.vtkActor() 932 self._actor.retrieve_object = weak_ref_to(self) 933 self._actor.SetMapper(self.mapper) 934 self.properties = self._actor.GetProperty() 935 936 self.name = "TetMesh" 937 938 # print('TetMesh inputtype', type(inputobj)) 939 940 ################### 941 if inputobj is None: 942 self.dataset = vtki.vtkUnstructuredGrid() 943 944 elif isinstance(inputobj, vtki.vtkUnstructuredGrid): 945 self.dataset = inputobj 946 947 elif isinstance(inputobj, UnstructuredGrid): 948 self.dataset = inputobj.dataset 949 950 elif "TetgenIO" in str(type(inputobj)): # tetgenpy object 951 inputobj = [inputobj.points(), inputobj.tetrahedra()] 952 953 elif isinstance(inputobj, vtki.vtkRectilinearGrid): 954 r2t = vtki.new("RectilinearGridToTetrahedra") 955 r2t.SetInputData(inputobj) 956 r2t.RememberVoxelIdOn() 957 r2t.SetTetraPerCellTo6() 958 r2t.Update() 959 self.dataset = r2t.GetOutput() 960 961 elif isinstance(inputobj, vtki.vtkDataSet): 962 r2t = vtki.new("DataSetTriangleFilter") 963 r2t.SetInputData(inputobj) 964 r2t.TetrahedraOnlyOn() 965 r2t.Update() 966 self.dataset = r2t.GetOutput() 967 968 elif isinstance(inputobj, str): 969 if "https://" in inputobj: 970 inputobj = download(inputobj, verbose=False) 971 if inputobj.endswith(".vtu"): 972 reader = vtki.new("XMLUnstructuredGridReader") 973 else: 974 reader = vtki.new("UnstructuredGridReader") 975 976 if not os.path.isfile(inputobj): 977 # for some reason vtk Reader does not complain 978 vedo.logger.error(f"file {inputobj} not found") 979 raise FileNotFoundError 980 981 self.filename = inputobj 982 reader.SetFileName(inputobj) 983 reader.Update() 984 ug = reader.GetOutput() 985 986 tt = vtki.new("DataSetTriangleFilter") 987 tt.SetInputData(ug) 988 tt.SetTetrahedraOnly(True) 989 tt.Update() 990 self.dataset = tt.GetOutput() 991 992 ############################### 993 if utils.is_sequence(inputobj): 994 self.dataset = vtki.vtkUnstructuredGrid() 995 996 points, cells = inputobj 997 if len(points) == 0: 998 return 999 if not utils.is_sequence(points[0]): 1000 return 1001 if len(cells) == 0: 1002 return 1003 1004 if not utils.is_sequence(cells[0]): 1005 tets = [] 1006 nf = cells[0] + 1 1007 for i, cl in enumerate(cells): 1008 if i in (nf, 0): 1009 k = i + 1 1010 nf = cl + k 1011 cell = [cells[j + k] for j in range(cl)] 1012 tets.append(cell) 1013 cells = tets 1014 1015 source_points = vtki.vtkPoints() 1016 varr = utils.numpy2vtk(points, dtype=np.float32) 1017 source_points.SetData(varr) 1018 self.dataset.SetPoints(source_points) 1019 1020 source_tets = vtki.vtkCellArray() 1021 for f in cells: 1022 ele = vtki.vtkTetra() 1023 pid = ele.GetPointIds() 1024 for i, fi in enumerate(f): 1025 pid.SetId(i, fi) 1026 source_tets.InsertNextCell(ele) 1027 self.dataset.SetCells(vtki.cell_types["TETRA"], source_tets) 1028 1029 if not self.dataset: 1030 vedo.logger.error(f"cannot understand input type {type(inputobj)}") 1031 return 1032 1033 self.properties.SetColor(0.352, 0.612, 0.996) # blue7 1034 1035 self.pipeline = utils.OperationNode( 1036 self, comment=f"#tets {self.dataset.GetNumberOfCells()}", c="#9e2a2b" 1037 ) 1038 1039 ################################################################## 1040 def __str__(self): 1041 """Print a string summary of the `TetMesh` object.""" 1042 module = self.__class__.__module__ 1043 name = self.__class__.__name__ 1044 out = vedo.printc( 1045 f"{module}.{name} at ({hex(self.memory_address())})".ljust(75), 1046 c="c", bold=True, invert=True, return_string=True, 1047 ) 1048 out += "\x1b[0m\u001b[36m" 1049 1050 out += "nr. of verts".ljust(14) + ": " + str(self.npoints) + "\n" 1051 out += "nr. of tetras".ljust(14) + ": " + str(self.ncells) + "\n" 1052 1053 if self.npoints: 1054 out+="size".ljust(14)+ ": average=" + utils.precision(self.average_size(),6) 1055 out+=", diagonal="+ utils.precision(self.diagonal_size(), 6)+ "\n" 1056 out+="center of mass".ljust(14) + ": " + utils.precision(self.center_of_mass(),6)+"\n" 1057 1058 bnds = self.bounds() 1059 bx1, bx2 = utils.precision(bnds[0], 3), utils.precision(bnds[1], 3) 1060 by1, by2 = utils.precision(bnds[2], 3), utils.precision(bnds[3], 3) 1061 bz1, bz2 = utils.precision(bnds[4], 3), utils.precision(bnds[5], 3) 1062 out += "bounds".ljust(14) + ":" 1063 out += " x=(" + bx1 + ", " + bx2 + ")," 1064 out += " y=(" + by1 + ", " + by2 + ")," 1065 out += " z=(" + bz1 + ", " + bz2 + ")\n" 1066 1067 for key in self.pointdata.keys(): 1068 arr = self.pointdata[key] 1069 rng = utils.precision(arr.min(), 3) + ", " + utils.precision(arr.max(), 3) 1070 mark_active = "pointdata" 1071 a_scalars = self.dataset.GetPointData().GetScalars() 1072 a_vectors = self.dataset.GetPointData().GetVectors() 1073 a_tensors = self.dataset.GetPointData().GetTensors() 1074 if a_scalars and a_scalars.GetName() == key: 1075 mark_active += " *" 1076 elif a_vectors and a_vectors.GetName() == key: 1077 mark_active += " **" 1078 elif a_tensors and a_tensors.GetName() == key: 1079 mark_active += " ***" 1080 out += mark_active.ljust(14) + f': "{key}" ({arr.dtype}), ndim={arr.ndim}' 1081 out += f", range=({rng})\n" 1082 1083 for key in self.celldata.keys(): 1084 arr = self.celldata[key] 1085 rng = utils.precision(arr.min(), 3) + ", " + utils.precision(arr.max(), 3) 1086 mark_active = "celldata" 1087 a_scalars = self.dataset.GetCellData().GetScalars() 1088 a_vectors = self.dataset.GetCellData().GetVectors() 1089 a_tensors = self.dataset.GetCellData().GetTensors() 1090 if a_scalars and a_scalars.GetName() == key: 1091 mark_active += " *" 1092 elif a_vectors and a_vectors.GetName() == key: 1093 mark_active += " **" 1094 elif a_tensors and a_tensors.GetName() == key: 1095 mark_active += " ***" 1096 out += mark_active.ljust(14) + f': "{key}" ({arr.dtype}), ndim={arr.ndim}' 1097 out += f", range=({rng})\n" 1098 1099 for key in self.metadata.keys(): 1100 arr = self.metadata[key] 1101 out += "metadata".ljust(14) + ": " + f'"{key}" ({len(arr)} values)\n' 1102 1103 return out.rstrip() + "\x1b[0m" 1104 1105 def _repr_html_(self): 1106 """ 1107 HTML representation of the TetMesh object for Jupyter Notebooks. 1108 1109 Returns: 1110 HTML text with the image and some properties. 1111 """ 1112 import io 1113 import base64 1114 from PIL import Image 1115 1116 library_name = "vedo.grids.TetMesh" 1117 help_url = "https://vedo.embl.es/docs/vedo/grids.html#TetMesh" 1118 1119 arr = self.thumbnail() 1120 im = Image.fromarray(arr) 1121 buffered = io.BytesIO() 1122 im.save(buffered, format="PNG", quality=100) 1123 encoded = base64.b64encode(buffered.getvalue()).decode("utf-8") 1124 url = "data:image/png;base64," + encoded 1125 image = f"<img src='{url}'></img>" 1126 1127 bounds = "<br/>".join( 1128 [ 1129 utils.precision(min_x, 4) + " ... " + utils.precision(max_x, 4) 1130 for min_x, max_x in zip(self.bounds()[::2], self.bounds()[1::2]) 1131 ] 1132 ) 1133 1134 help_text = "" 1135 if self.name: 1136 help_text += f"<b> {self.name}:   </b>" 1137 help_text += '<b><a href="' + help_url + '" target="_blank">' + library_name + "</a></b>" 1138 if self.filename: 1139 dots = "" 1140 if len(self.filename) > 30: 1141 dots = "..." 1142 help_text += f"<br/><code><i>({dots}{self.filename[-30:]})</i></code>" 1143 1144 pdata = "" 1145 if self.dataset.GetPointData().GetScalars(): 1146 if self.dataset.GetPointData().GetScalars().GetName(): 1147 name = self.dataset.GetPointData().GetScalars().GetName() 1148 pdata = "<tr><td><b> point data array </b></td><td>" + name + "</td></tr>" 1149 1150 cdata = "" 1151 if self.dataset.GetCellData().GetScalars(): 1152 if self.dataset.GetCellData().GetScalars().GetName(): 1153 name = self.dataset.GetCellData().GetScalars().GetName() 1154 cdata = "<tr><td><b> cell data array </b></td><td>" + name + "</td></tr>" 1155 1156 pts = self.coordinates 1157 cm = np.mean(pts, axis=0) 1158 1159 allt = [ 1160 "<table>", 1161 "<tr>", 1162 "<td>", image, "</td>", 1163 "<td style='text-align: center; vertical-align: center;'><br/>", help_text, 1164 "<table>", 1165 "<tr><td><b> bounds </b> <br/> (x/y/z) </td><td>" + str(bounds) + "</td></tr>", 1166 "<tr><td><b> center of mass </b></td><td>" + utils.precision(cm,3) + "</td></tr>", 1167 "<tr><td><b> nr. points / tets </b></td><td>" 1168 + str(self.npoints) + " / " + str(self.ncells) + "</td></tr>", 1169 pdata, 1170 cdata, 1171 "</table>", 1172 "</table>", 1173 ] 1174 return "\n".join(allt) 1175 1176 def compute_quality(self, metric=7) -> np.ndarray: 1177 """ 1178 Calculate functions of quality for the elements of a tetrahedral mesh. 1179 This method adds to the mesh a cell array named "Quality". 1180 1181 Arguments: 1182 metric : (int) 1183 type of estimators: 1184 - EDGE RATIO, 0 1185 - ASPECT RATIO, 1 1186 - RADIUS RATIO, 2 1187 - ASPECT FROBENIUS, 3 1188 - MIN_ANGLE, 4 1189 - COLLAPSE RATIO, 5 1190 - ASPECT GAMMA, 6 1191 - VOLUME, 7 1192 1193 See class [vtkMeshQuality](https://vtk.org/doc/nightly/html/classvtkMeshQuality.html) 1194 for an explanation of the meaning of each metric.. 1195 """ 1196 qf = vtki.new("MeshQuality") 1197 qf.SetInputData(self.dataset) 1198 qf.SetTetQualityMeasure(metric) 1199 qf.SaveCellQualityOn() 1200 qf.Update() 1201 self._update(qf.GetOutput()) 1202 return utils.vtk2numpy(qf.GetOutput().GetCellData().GetArray("Quality")) 1203 1204 def check_validity(self, tol=0) -> np.ndarray: 1205 """ 1206 Return an array of possible problematic tets following this convention: 1207 ```python 1208 Valid = 0 1209 WrongNumberOfPoints = 01 1210 IntersectingEdges = 02 1211 IntersectingFaces = 04 1212 NoncontiguousEdges = 08 1213 Nonconvex = 10 1214 OrientedIncorrectly = 20 1215 ``` 1216 1217 Arguments: 1218 tol : (float) 1219 This value is used as an epsilon for floating point 1220 equality checks throughout the cell checking process. 1221 """ 1222 vald = vtki.new("CellValidator") 1223 if tol: 1224 vald.SetTolerance(tol) 1225 vald.SetInputData(self.dataset) 1226 vald.Update() 1227 varr = vald.GetOutput().GetCellData().GetArray("ValidityState") 1228 return utils.vtk2numpy(varr) 1229 1230 def decimate(self, scalars_name: str, fraction=0.5, n=0) -> Self: 1231 """ 1232 Downsample the number of tets in a TetMesh to a specified fraction. 1233 Either `fraction` or `n` must be set. 1234 1235 Arguments: 1236 fraction : (float) 1237 the desired final fraction of the total. 1238 n : (int) 1239 the desired number of final tets 1240 1241 .. note:: setting `fraction=0.1` leaves 10% of the original nr of tets. 1242 """ 1243 decimate = vtki.new("UnstructuredGridQuadricDecimation") 1244 decimate.SetInputData(self.dataset) 1245 decimate.SetScalarsName(scalars_name) 1246 1247 if n: # n = desired number of points 1248 decimate.SetNumberOfTetsOutput(n) 1249 else: 1250 decimate.SetTargetReduction(1 - fraction) 1251 decimate.Update() 1252 self._update(decimate.GetOutput()) 1253 self.pipeline = utils.OperationNode( 1254 "decimate", comment=f"array: {scalars_name}", c="#edabab", parents=[self] 1255 ) 1256 return self 1257 1258 def subdivide(self) -> Self: 1259 """ 1260 Increase the number of tetrahedrons of a `TetMesh`. 1261 Subdivides each tetrahedron into twelve smaller tetras. 1262 """ 1263 sd = vtki.new("SubdivideTetra") 1264 sd.SetInputData(self.dataset) 1265 sd.Update() 1266 self._update(sd.GetOutput()) 1267 self.pipeline = utils.OperationNode("subdivide", c="#edabab", parents=[self]) 1268 return self 1269 1270 def generate_random_points(self, n, min_radius=0) -> "vedo.Points": 1271 """ 1272 Generate `n` uniformly distributed random points 1273 inside the tetrahedral mesh. 1274 1275 A new point data array is added to the output points 1276 called "OriginalCellID" which contains the index of 1277 the cell ID in which the point was generated. 1278 1279 Arguments: 1280 n : (int) 1281 number of points to generate. 1282 min_radius: (float) 1283 impose a minimum distance between points. 1284 If `min_radius` is set to 0, the points are 1285 generated uniformly at random inside the mesh. 1286 If `min_radius` is set to a positive value, 1287 the points are generated uniformly at random 1288 inside the mesh, but points closer than `min_radius` 1289 to any other point are discarded. 1290 1291 Returns a `vedo.Points` object. 1292 1293 Note: 1294 Consider using `points.probe(msh)` to interpolate 1295 any existing mesh data onto the points. 1296 1297 Example: 1298 ```python 1299 from vedo import * 1300 tmesh = TetMesh(dataurl + "limb.vtu").alpha(0.2) 1301 pts = tmesh.generate_random_points(20000, min_radius=10) 1302 print(pts.pointdata["OriginalCellID"]) 1303 show(pts, tmesh, axes=1).close() 1304 ``` 1305 """ 1306 cmesh = self.compute_cell_size() 1307 tets = cmesh.cells 1308 verts = cmesh.coordinates 1309 cumul = np.cumsum(np.abs(cmesh.celldata["Volume"])) 1310 1311 out_pts = [] 1312 orig_cell = [] 1313 for _ in range(n): 1314 random_area = np.random.random() * cumul[-1] 1315 it = np.searchsorted(cumul, random_area) 1316 A, B, C, D = verts[tets[it]] 1317 r1, r2, r3 = sorted(np.random.random(3)) 1318 p = r1 * A + (r2 - r1) * B + (r3 - r2) * C + (1 - r3) * D 1319 out_pts.append(p) 1320 orig_cell.append(it) 1321 orig_cellnp = np.array(orig_cell, dtype=np.uint32) 1322 1323 vpts = vedo.pointcloud.Points(out_pts) 1324 vpts.pointdata["OriginalCellID"] = orig_cellnp 1325 1326 if min_radius > 0: 1327 vpts.subsample(min_radius, absolute=True) 1328 1329 vpts.point_size(5).color("k1") 1330 vpts.name = "RandomPoints" 1331 vpts.pipeline = utils.OperationNode( 1332 "generate_random_points", c="#edabab", parents=[self]) 1333 return vpts 1334 1335 def isosurface(self, value=True, flying_edges=None) -> "vedo.Mesh": 1336 """ 1337 Return a `vedo.Mesh` isosurface. 1338 The "isosurface" is the surface of the region of points 1339 whose values equal to `value`. 1340 1341 Set `value` to a single value or list of values to compute the isosurface(s). 1342 1343 Note that flying_edges option is not available for `TetMesh`. 1344 """ 1345 if flying_edges is not None: 1346 vedo.logger.warning("flying_edges option is not available for TetMesh.") 1347 1348 if not self.dataset.GetPointData().GetScalars(): 1349 vedo.logger.warning( 1350 "in isosurface() no scalar pointdata found. " 1351 "Mappping cells to points." 1352 ) 1353 self.map_cells_to_points() 1354 scrange = self.dataset.GetPointData().GetScalars().GetRange() 1355 cf = vtki.new("ContourFilter") # vtki.new("ContourGrid") 1356 cf.SetInputData(self.dataset) 1357 1358 if utils.is_sequence(value): 1359 cf.SetNumberOfContours(len(value)) 1360 for i, t in enumerate(value): 1361 cf.SetValue(i, t) 1362 cf.Update() 1363 else: 1364 if value is True: 1365 value = (2 * scrange[0] + scrange[1]) / 3.0 1366 cf.SetValue(0, value) 1367 cf.Update() 1368 1369 msh = Mesh(cf.GetOutput(), c=None) 1370 msh.copy_properties_from(self) 1371 msh.pipeline = utils.OperationNode("isosurface", c="#edabab", parents=[self]) 1372 return msh 1373 1374 def slice(self, origin=(0, 0, 0), normal=(1, 0, 0)) -> "vedo.Mesh": 1375 """ 1376 Return a 2D slice of the mesh by a plane passing through origin and assigned normal. 1377 """ 1378 strn = str(normal) 1379 if strn == "x": normal = (1, 0, 0) 1380 elif strn == "y": normal = (0, 1, 0) 1381 elif strn == "z": normal = (0, 0, 1) 1382 elif strn == "-x": normal = (-1, 0, 0) 1383 elif strn == "-y": normal = (0, -1, 0) 1384 elif strn == "-z": normal = (0, 0, -1) 1385 plane = vtki.new("Plane") 1386 plane.SetOrigin(origin) 1387 plane.SetNormal(normal) 1388 1389 cc = vtki.new("Cutter") 1390 cc.SetInputData(self.dataset) 1391 cc.SetCutFunction(plane) 1392 cc.Update() 1393 msh = Mesh(cc.GetOutput()).flat().lighting("ambient") 1394 msh.copy_properties_from(self) 1395 msh.pipeline = utils.OperationNode("slice", c="#edabab", parents=[self]) 1396 return msh
The class describing tetrahedral meshes.
920 def __init__(self, inputobj=None): 921 """ 922 Arguments: 923 inputobj : (vtkUnstructuredGrid, list, str, tetgenpy.TetgenIO) 924 list of points and tet indices, or filename 925 """ 926 super().__init__() 927 928 self.dataset = None 929 930 self.mapper = vtki.new("PolyDataMapper") 931 self._actor = vtki.vtkActor() 932 self._actor.retrieve_object = weak_ref_to(self) 933 self._actor.SetMapper(self.mapper) 934 self.properties = self._actor.GetProperty() 935 936 self.name = "TetMesh" 937 938 # print('TetMesh inputtype', type(inputobj)) 939 940 ################### 941 if inputobj is None: 942 self.dataset = vtki.vtkUnstructuredGrid() 943 944 elif isinstance(inputobj, vtki.vtkUnstructuredGrid): 945 self.dataset = inputobj 946 947 elif isinstance(inputobj, UnstructuredGrid): 948 self.dataset = inputobj.dataset 949 950 elif "TetgenIO" in str(type(inputobj)): # tetgenpy object 951 inputobj = [inputobj.points(), inputobj.tetrahedra()] 952 953 elif isinstance(inputobj, vtki.vtkRectilinearGrid): 954 r2t = vtki.new("RectilinearGridToTetrahedra") 955 r2t.SetInputData(inputobj) 956 r2t.RememberVoxelIdOn() 957 r2t.SetTetraPerCellTo6() 958 r2t.Update() 959 self.dataset = r2t.GetOutput() 960 961 elif isinstance(inputobj, vtki.vtkDataSet): 962 r2t = vtki.new("DataSetTriangleFilter") 963 r2t.SetInputData(inputobj) 964 r2t.TetrahedraOnlyOn() 965 r2t.Update() 966 self.dataset = r2t.GetOutput() 967 968 elif isinstance(inputobj, str): 969 if "https://" in inputobj: 970 inputobj = download(inputobj, verbose=False) 971 if inputobj.endswith(".vtu"): 972 reader = vtki.new("XMLUnstructuredGridReader") 973 else: 974 reader = vtki.new("UnstructuredGridReader") 975 976 if not os.path.isfile(inputobj): 977 # for some reason vtk Reader does not complain 978 vedo.logger.error(f"file {inputobj} not found") 979 raise FileNotFoundError 980 981 self.filename = inputobj 982 reader.SetFileName(inputobj) 983 reader.Update() 984 ug = reader.GetOutput() 985 986 tt = vtki.new("DataSetTriangleFilter") 987 tt.SetInputData(ug) 988 tt.SetTetrahedraOnly(True) 989 tt.Update() 990 self.dataset = tt.GetOutput() 991 992 ############################### 993 if utils.is_sequence(inputobj): 994 self.dataset = vtki.vtkUnstructuredGrid() 995 996 points, cells = inputobj 997 if len(points) == 0: 998 return 999 if not utils.is_sequence(points[0]): 1000 return 1001 if len(cells) == 0: 1002 return 1003 1004 if not utils.is_sequence(cells[0]): 1005 tets = [] 1006 nf = cells[0] + 1 1007 for i, cl in enumerate(cells): 1008 if i in (nf, 0): 1009 k = i + 1 1010 nf = cl + k 1011 cell = [cells[j + k] for j in range(cl)] 1012 tets.append(cell) 1013 cells = tets 1014 1015 source_points = vtki.vtkPoints() 1016 varr = utils.numpy2vtk(points, dtype=np.float32) 1017 source_points.SetData(varr) 1018 self.dataset.SetPoints(source_points) 1019 1020 source_tets = vtki.vtkCellArray() 1021 for f in cells: 1022 ele = vtki.vtkTetra() 1023 pid = ele.GetPointIds() 1024 for i, fi in enumerate(f): 1025 pid.SetId(i, fi) 1026 source_tets.InsertNextCell(ele) 1027 self.dataset.SetCells(vtki.cell_types["TETRA"], source_tets) 1028 1029 if not self.dataset: 1030 vedo.logger.error(f"cannot understand input type {type(inputobj)}") 1031 return 1032 1033 self.properties.SetColor(0.352, 0.612, 0.996) # blue7 1034 1035 self.pipeline = utils.OperationNode( 1036 self, comment=f"#tets {self.dataset.GetNumberOfCells()}", c="#9e2a2b" 1037 )
Arguments:
- inputobj : (vtkUnstructuredGrid, list, str, tetgenpy.TetgenIO) list of points and tet indices, or filename
1176 def compute_quality(self, metric=7) -> np.ndarray: 1177 """ 1178 Calculate functions of quality for the elements of a tetrahedral mesh. 1179 This method adds to the mesh a cell array named "Quality". 1180 1181 Arguments: 1182 metric : (int) 1183 type of estimators: 1184 - EDGE RATIO, 0 1185 - ASPECT RATIO, 1 1186 - RADIUS RATIO, 2 1187 - ASPECT FROBENIUS, 3 1188 - MIN_ANGLE, 4 1189 - COLLAPSE RATIO, 5 1190 - ASPECT GAMMA, 6 1191 - VOLUME, 7 1192 1193 See class [vtkMeshQuality](https://vtk.org/doc/nightly/html/classvtkMeshQuality.html) 1194 for an explanation of the meaning of each metric.. 1195 """ 1196 qf = vtki.new("MeshQuality") 1197 qf.SetInputData(self.dataset) 1198 qf.SetTetQualityMeasure(metric) 1199 qf.SaveCellQualityOn() 1200 qf.Update() 1201 self._update(qf.GetOutput()) 1202 return utils.vtk2numpy(qf.GetOutput().GetCellData().GetArray("Quality"))
Calculate functions of quality for the elements of a tetrahedral mesh. This method adds to the mesh a cell array named "Quality".
Arguments:
- metric : (int)
type of estimators:
- EDGE RATIO, 0
- ASPECT RATIO, 1
- RADIUS RATIO, 2
- ASPECT FROBENIUS, 3
- MIN_ANGLE, 4
- COLLAPSE RATIO, 5
- ASPECT GAMMA, 6
- VOLUME, 7
See class vtkMeshQuality for an explanation of the meaning of each metric..
1204 def check_validity(self, tol=0) -> np.ndarray: 1205 """ 1206 Return an array of possible problematic tets following this convention: 1207 ```python 1208 Valid = 0 1209 WrongNumberOfPoints = 01 1210 IntersectingEdges = 02 1211 IntersectingFaces = 04 1212 NoncontiguousEdges = 08 1213 Nonconvex = 10 1214 OrientedIncorrectly = 20 1215 ``` 1216 1217 Arguments: 1218 tol : (float) 1219 This value is used as an epsilon for floating point 1220 equality checks throughout the cell checking process. 1221 """ 1222 vald = vtki.new("CellValidator") 1223 if tol: 1224 vald.SetTolerance(tol) 1225 vald.SetInputData(self.dataset) 1226 vald.Update() 1227 varr = vald.GetOutput().GetCellData().GetArray("ValidityState") 1228 return utils.vtk2numpy(varr)
Return an array of possible problematic tets following this convention:
Valid = 0
WrongNumberOfPoints = 01
IntersectingEdges = 02
IntersectingFaces = 04
NoncontiguousEdges = 08
Nonconvex = 10
OrientedIncorrectly = 20
Arguments:
- tol : (float) This value is used as an epsilon for floating point equality checks throughout the cell checking process.
1230 def decimate(self, scalars_name: str, fraction=0.5, n=0) -> Self: 1231 """ 1232 Downsample the number of tets in a TetMesh to a specified fraction. 1233 Either `fraction` or `n` must be set. 1234 1235 Arguments: 1236 fraction : (float) 1237 the desired final fraction of the total. 1238 n : (int) 1239 the desired number of final tets 1240 1241 .. note:: setting `fraction=0.1` leaves 10% of the original nr of tets. 1242 """ 1243 decimate = vtki.new("UnstructuredGridQuadricDecimation") 1244 decimate.SetInputData(self.dataset) 1245 decimate.SetScalarsName(scalars_name) 1246 1247 if n: # n = desired number of points 1248 decimate.SetNumberOfTetsOutput(n) 1249 else: 1250 decimate.SetTargetReduction(1 - fraction) 1251 decimate.Update() 1252 self._update(decimate.GetOutput()) 1253 self.pipeline = utils.OperationNode( 1254 "decimate", comment=f"array: {scalars_name}", c="#edabab", parents=[self] 1255 ) 1256 return self
Downsample the number of tets in a TetMesh to a specified fraction.
Either fraction or n must be set.
Arguments:
- fraction : (float) the desired final fraction of the total.
- n : (int) the desired number of final tets
setting fraction=0.1 leaves 10% of the original nr of tets.
1258 def subdivide(self) -> Self: 1259 """ 1260 Increase the number of tetrahedrons of a `TetMesh`. 1261 Subdivides each tetrahedron into twelve smaller tetras. 1262 """ 1263 sd = vtki.new("SubdivideTetra") 1264 sd.SetInputData(self.dataset) 1265 sd.Update() 1266 self._update(sd.GetOutput()) 1267 self.pipeline = utils.OperationNode("subdivide", c="#edabab", parents=[self]) 1268 return self
Increase the number of tetrahedrons of a TetMesh.
Subdivides each tetrahedron into twelve smaller tetras.
1270 def generate_random_points(self, n, min_radius=0) -> "vedo.Points": 1271 """ 1272 Generate `n` uniformly distributed random points 1273 inside the tetrahedral mesh. 1274 1275 A new point data array is added to the output points 1276 called "OriginalCellID" which contains the index of 1277 the cell ID in which the point was generated. 1278 1279 Arguments: 1280 n : (int) 1281 number of points to generate. 1282 min_radius: (float) 1283 impose a minimum distance between points. 1284 If `min_radius` is set to 0, the points are 1285 generated uniformly at random inside the mesh. 1286 If `min_radius` is set to a positive value, 1287 the points are generated uniformly at random 1288 inside the mesh, but points closer than `min_radius` 1289 to any other point are discarded. 1290 1291 Returns a `vedo.Points` object. 1292 1293 Note: 1294 Consider using `points.probe(msh)` to interpolate 1295 any existing mesh data onto the points. 1296 1297 Example: 1298 ```python 1299 from vedo import * 1300 tmesh = TetMesh(dataurl + "limb.vtu").alpha(0.2) 1301 pts = tmesh.generate_random_points(20000, min_radius=10) 1302 print(pts.pointdata["OriginalCellID"]) 1303 show(pts, tmesh, axes=1).close() 1304 ``` 1305 """ 1306 cmesh = self.compute_cell_size() 1307 tets = cmesh.cells 1308 verts = cmesh.coordinates 1309 cumul = np.cumsum(np.abs(cmesh.celldata["Volume"])) 1310 1311 out_pts = [] 1312 orig_cell = [] 1313 for _ in range(n): 1314 random_area = np.random.random() * cumul[-1] 1315 it = np.searchsorted(cumul, random_area) 1316 A, B, C, D = verts[tets[it]] 1317 r1, r2, r3 = sorted(np.random.random(3)) 1318 p = r1 * A + (r2 - r1) * B + (r3 - r2) * C + (1 - r3) * D 1319 out_pts.append(p) 1320 orig_cell.append(it) 1321 orig_cellnp = np.array(orig_cell, dtype=np.uint32) 1322 1323 vpts = vedo.pointcloud.Points(out_pts) 1324 vpts.pointdata["OriginalCellID"] = orig_cellnp 1325 1326 if min_radius > 0: 1327 vpts.subsample(min_radius, absolute=True) 1328 1329 vpts.point_size(5).color("k1") 1330 vpts.name = "RandomPoints" 1331 vpts.pipeline = utils.OperationNode( 1332 "generate_random_points", c="#edabab", parents=[self]) 1333 return vpts
Generate n uniformly distributed random points
inside the tetrahedral mesh.
A new point data array is added to the output points called "OriginalCellID" which contains the index of the cell ID in which the point was generated.
Arguments:
- n : (int) number of points to generate.
- min_radius: (float)
impose a minimum distance between points.
If
min_radiusis set to 0, the points are generated uniformly at random inside the mesh. Ifmin_radiusis set to a positive value, the points are generated uniformly at random inside the mesh, but points closer thanmin_radiusto any other point are discarded.
Returns a vedo.Points object.
Note:
Consider using
points.probe(msh)to interpolate any existing mesh data onto the points.
Example:
from vedo import *
tmesh = TetMesh(dataurl + "limb.vtu").alpha(0.2)
pts = tmesh.generate_random_points(20000, min_radius=10)
print(pts.pointdata["OriginalCellID"])
show(pts, tmesh, axes=1).close()
1335 def isosurface(self, value=True, flying_edges=None) -> "vedo.Mesh": 1336 """ 1337 Return a `vedo.Mesh` isosurface. 1338 The "isosurface" is the surface of the region of points 1339 whose values equal to `value`. 1340 1341 Set `value` to a single value or list of values to compute the isosurface(s). 1342 1343 Note that flying_edges option is not available for `TetMesh`. 1344 """ 1345 if flying_edges is not None: 1346 vedo.logger.warning("flying_edges option is not available for TetMesh.") 1347 1348 if not self.dataset.GetPointData().GetScalars(): 1349 vedo.logger.warning( 1350 "in isosurface() no scalar pointdata found. " 1351 "Mappping cells to points." 1352 ) 1353 self.map_cells_to_points() 1354 scrange = self.dataset.GetPointData().GetScalars().GetRange() 1355 cf = vtki.new("ContourFilter") # vtki.new("ContourGrid") 1356 cf.SetInputData(self.dataset) 1357 1358 if utils.is_sequence(value): 1359 cf.SetNumberOfContours(len(value)) 1360 for i, t in enumerate(value): 1361 cf.SetValue(i, t) 1362 cf.Update() 1363 else: 1364 if value is True: 1365 value = (2 * scrange[0] + scrange[1]) / 3.0 1366 cf.SetValue(0, value) 1367 cf.Update() 1368 1369 msh = Mesh(cf.GetOutput(), c=None) 1370 msh.copy_properties_from(self) 1371 msh.pipeline = utils.OperationNode("isosurface", c="#edabab", parents=[self]) 1372 return msh
Return a vedo.Mesh isosurface.
The "isosurface" is the surface of the region of points
whose values equal to value.
Set value to a single value or list of values to compute the isosurface(s).
Note that flying_edges option is not available for TetMesh.
1374 def slice(self, origin=(0, 0, 0), normal=(1, 0, 0)) -> "vedo.Mesh": 1375 """ 1376 Return a 2D slice of the mesh by a plane passing through origin and assigned normal. 1377 """ 1378 strn = str(normal) 1379 if strn == "x": normal = (1, 0, 0) 1380 elif strn == "y": normal = (0, 1, 0) 1381 elif strn == "z": normal = (0, 0, 1) 1382 elif strn == "-x": normal = (-1, 0, 0) 1383 elif strn == "-y": normal = (0, -1, 0) 1384 elif strn == "-z": normal = (0, 0, -1) 1385 plane = vtki.new("Plane") 1386 plane.SetOrigin(origin) 1387 plane.SetNormal(normal) 1388 1389 cc = vtki.new("Cutter") 1390 cc.SetInputData(self.dataset) 1391 cc.SetCutFunction(plane) 1392 cc.Update() 1393 msh = Mesh(cc.GetOutput()).flat().lighting("ambient") 1394 msh.copy_properties_from(self) 1395 msh.pipeline = utils.OperationNode("slice", c="#edabab", parents=[self]) 1396 return msh
Return a 2D slice of the mesh by a plane passing through origin and assigned normal.
Inherited Members
- UnstructuredGrid
- actor
- merge
- copy
- clone
- bounds
- threshold
- shrink
- tomesh
- cell_types_array
- extract_cells_by_type
- extract_cells_by_id
- find_cell
- clean
- extract_cells_on_plane
- extract_cells_on_sphere
- extract_cells_on_cylinder
- cut_with_plane
- cut_with_box
- cut_with_mesh
- vedo.core.PointAlgorithms
- apply_transform
- apply_transform_from_actor
- pos
- shift
- x
- y
- z
- rotate
- rotate_x
- rotate_y
- rotate_z
- reorient
- scale
- vedo.core.CommonAlgorithms
- pointdata
- celldata
- metadata
- rename
- memory_address
- memory_size
- modified
- box
- update_dataset
- xbounds
- ybounds
- zbounds
- diagonal_size
- average_size
- center_of_mass
- copy_data_from
- inputdata
- npoints
- nvertices
- ncells
- cell_centers
- lines
- lines_as_flat_array
- mark_boundaries
- find_cells_in_bounds
- find_cells_along_line
- find_cells_along_plane
- keep_cell_types
- map_cells_to_points
- vertices
- points
- coordinates
- cells_as_flat_array
- cells
- cell_edge_neighbors
- map_points_to_cells
- resample_data_from
- interpolate_data_from
- add_ids
- gradient
- divergence
- vorticity
- probe
- compute_cell_size
- generate_random_data
- integrate_data
- write
- signed_distance
- unsigned_distance
- smooth_data
- compute_streamlines
- vedo.visual.MeshVisual
- follow_camera
- wireframe
- flat
- phong
- backface_culling
- render_lines_as_tubes
- frontface_culling
- backcolor
- bc
- linewidth
- lw
- linecolor
- lc
- texture
- vedo.visual.PointsVisual
- clone2d
- copy_properties_from
- color
- c
- alpha
- lut_color_at
- opacity
- force_opaque
- force_translucent
- point_size
- ps
- render_points_as_spheres
- lighting
- point_blurring
- cellcolors
- pointcolors
- cmap
- add_trail
- update_trail
- add_shadow
- update_shadows
- labels
- labels2d
- legend
- flagpole
- flagpost
- caption
1400class RectilinearGrid(PointAlgorithms, MeshVisual): 1401 """ 1402 Build a rectilinear grid. 1403 """ 1404 1405 def __init__(self, inputobj=None): 1406 """ 1407 A RectilinearGrid is a dataset where edges are parallel to the coordinate axes. 1408 It can be thought of as a tessellation of a box in 3D space, similar to a `Volume` 1409 except that the cells are not necessarily cubes, but they can have different lengths 1410 along each axis. 1411 This can be useful to describe a volume with variable resolution where one needs 1412 to represent a region with higher detail with respect to another region. 1413 1414 Arguments: 1415 inputobj : (vtkRectilinearGrid, list, str) 1416 list of points and indices, or filename 1417 1418 Example: 1419 ```python 1420 from vedo import RectilinearGrid, show 1421 1422 xcoords = 7 + np.sqrt(np.arange(0,2500,25)) 1423 ycoords = np.arange(0, 20) 1424 zcoords = np.arange(0, 20) 1425 1426 rgrid = RectilinearGrid([xcoords, ycoords, zcoords]) 1427 1428 print(rgrid) 1429 print(rgrid.x_coordinates().shape) 1430 print(rgrid.compute_structured_coords([20,10,11])) 1431 1432 msh = rgrid.tomesh().lw(1) 1433 1434 show(msh, axes=1, viewup="z") 1435 ``` 1436 """ 1437 1438 super().__init__() 1439 1440 self.dataset = None 1441 1442 self.mapper = vtki.new("PolyDataMapper") 1443 self._actor = vtki.vtkActor() 1444 self._actor.retrieve_object = weak_ref_to(self) 1445 self._actor.SetMapper(self.mapper) 1446 self.properties = self._actor.GetProperty() 1447 1448 self.transform = LinearTransform() 1449 self.point_locator = None 1450 self.cell_locator = None 1451 self.line_locator = None 1452 1453 self.name = "RectilinearGrid" 1454 self.filename = "" 1455 1456 self.info = {} 1457 self.time = time.time() 1458 1459 ############################### 1460 if inputobj is None: 1461 self.dataset = vtki.vtkRectilinearGrid() 1462 1463 elif isinstance(inputobj, vtki.vtkRectilinearGrid): 1464 self.dataset = inputobj 1465 1466 elif isinstance(inputobj, RectilinearGrid): 1467 self.dataset = inputobj.dataset 1468 1469 elif isinstance(inputobj, str): 1470 if "https://" in inputobj: 1471 inputobj = download(inputobj, verbose=False) 1472 if inputobj.endswith(".vtr"): 1473 reader = vtki.new("XMLRectilinearGridReader") 1474 else: 1475 reader = vtki.new("RectilinearGridReader") 1476 self.filename = inputobj 1477 reader.SetFileName(inputobj) 1478 reader.Update() 1479 self.dataset = reader.GetOutput() 1480 1481 elif utils.is_sequence(inputobj): 1482 self.dataset = vtki.vtkRectilinearGrid() 1483 xcoords, ycoords, zcoords = inputobj 1484 nx, ny, nz = len(xcoords), len(ycoords), len(zcoords) 1485 self.dataset.SetDimensions(nx, ny, nz) 1486 self.dataset.SetXCoordinates(utils.numpy2vtk(xcoords)) 1487 self.dataset.SetYCoordinates(utils.numpy2vtk(ycoords)) 1488 self.dataset.SetZCoordinates(utils.numpy2vtk(zcoords)) 1489 1490 ############################### 1491 1492 if not self.dataset: 1493 vedo.logger.error(f"RectilinearGrid: cannot understand input type {type(inputobj)}") 1494 return 1495 1496 self.properties.SetColor(0.352, 0.612, 0.996) # blue7 1497 1498 self.pipeline = utils.OperationNode( 1499 self, comment=f"#tets {self.dataset.GetNumberOfCells()}", c="#9e2a2b" 1500 ) 1501 1502 @property 1503 def actor(self): 1504 """Return the `vtkActor` of the object.""" 1505 gf = vtki.new("GeometryFilter") 1506 gf.SetInputData(self.dataset) 1507 gf.Update() 1508 self.mapper.SetInputData(gf.GetOutput()) 1509 self.mapper.Modified() 1510 return self._actor 1511 1512 @actor.setter 1513 def actor(self, _): 1514 pass 1515 1516 def _update(self, data, reset_locators=False): 1517 self.dataset = data 1518 if reset_locators: 1519 self.cell_locator = None 1520 self.point_locator = None 1521 return self 1522 1523 ################################################################## 1524 def __str__(self): 1525 """Print a summary for the `RectilinearGrid` object.""" 1526 module = self.__class__.__module__ 1527 name = self.__class__.__name__ 1528 out = vedo.printc( 1529 f"{module}.{name} at ({hex(self.memory_address())})".ljust(75), 1530 c="c", bold=True, invert=True, return_string=True, 1531 ) 1532 out += "\x1b[0m\x1b[36;1m" 1533 1534 out += "name".ljust(14) + ": " + str(self.name) + "\n" 1535 if self.filename: 1536 out += "filename".ljust(14) + ": " + str(self.filename) + "\n" 1537 1538 out += "dimensions".ljust(14) + ": " + str(self.dataset.GetDimensions()) + "\n" 1539 1540 out += "center".ljust(14) + ": " 1541 out += utils.precision(self.dataset.GetCenter(), 6) + "\n" 1542 1543 bnds = self.bounds() 1544 bx1, bx2 = utils.precision(bnds[0], 3), utils.precision(bnds[1], 3) 1545 by1, by2 = utils.precision(bnds[2], 3), utils.precision(bnds[3], 3) 1546 bz1, bz2 = utils.precision(bnds[4], 3), utils.precision(bnds[5], 3) 1547 out += "bounds".ljust(14) + ":" 1548 out += " x=(" + bx1 + ", " + bx2 + ")," 1549 out += " y=(" + by1 + ", " + by2 + ")," 1550 out += " z=(" + bz1 + ", " + bz2 + ")\n" 1551 1552 out += "memory size".ljust(14) + ": " 1553 out += utils.precision(self.dataset.GetActualMemorySize() / 1024, 2) + " MB\n" 1554 1555 for key in self.pointdata.keys(): 1556 arr = self.pointdata[key] 1557 rng = utils.precision(arr.min(), 3) + ", " + utils.precision(arr.max(), 3) 1558 mark_active = "pointdata" 1559 a_scalars = self.dataset.GetPointData().GetScalars() 1560 a_vectors = self.dataset.GetPointData().GetVectors() 1561 a_tensors = self.dataset.GetPointData().GetTensors() 1562 if a_scalars and a_scalars.GetName() == key: 1563 mark_active += " *" 1564 elif a_vectors and a_vectors.GetName() == key: 1565 mark_active += " **" 1566 elif a_tensors and a_tensors.GetName() == key: 1567 mark_active += " ***" 1568 out += mark_active.ljust(14) + f': "{key}" ({arr.dtype}), ndim={arr.ndim}' 1569 out += f", range=({rng})\n" 1570 1571 for key in self.celldata.keys(): 1572 arr = self.celldata[key] 1573 rng = utils.precision(arr.min(), 3) + ", " + utils.precision(arr.max(), 3) 1574 mark_active = "celldata" 1575 a_scalars = self.dataset.GetCellData().GetScalars() 1576 a_vectors = self.dataset.GetCellData().GetVectors() 1577 a_tensors = self.dataset.GetCellData().GetTensors() 1578 if a_scalars and a_scalars.GetName() == key: 1579 mark_active += " *" 1580 elif a_vectors and a_vectors.GetName() == key: 1581 mark_active += " **" 1582 elif a_tensors and a_tensors.GetName() == key: 1583 mark_active += " ***" 1584 out += mark_active.ljust(14) + f': "{key}" ({arr.dtype}), ndim={arr.ndim}' 1585 out += f", range=({rng})\n" 1586 1587 for key in self.metadata.keys(): 1588 arr = self.metadata[key] 1589 out += "metadata".ljust(14) + ": " + f'"{key}" ({len(arr)} values)\n' 1590 1591 return out.rstrip() + "\x1b[0m" 1592 1593 def _repr_html_(self): 1594 """ 1595 HTML representation of the RectilinearGrid object for Jupyter Notebooks. 1596 1597 Returns: 1598 HTML text with the image and some properties. 1599 """ 1600 import io 1601 import base64 1602 from PIL import Image 1603 1604 library_name = "vedo.grids.RectilinearGrid" 1605 help_url = "https://vedo.embl.es/docs/vedo/grids.html#RectilinearGrid" 1606 1607 m = self.tomesh().linewidth(1).lighting("off") 1608 arr= m.thumbnail(zoom=1, elevation=-30, azimuth=-30) 1609 1610 im = Image.fromarray(arr) 1611 buffered = io.BytesIO() 1612 im.save(buffered, format="PNG", quality=100) 1613 encoded = base64.b64encode(buffered.getvalue()).decode("utf-8") 1614 url = "data:image/png;base64," + encoded 1615 image = f"<img src='{url}'></img>" 1616 1617 bounds = "<br/>".join( 1618 [ 1619 utils.precision(min_x, 4) + " ... " + utils.precision(max_x, 4) 1620 for min_x, max_x in zip(self.bounds()[::2], self.bounds()[1::2]) 1621 ] 1622 ) 1623 1624 help_text = "" 1625 if self.name: 1626 help_text += f"<b> {self.name}:   </b>" 1627 help_text += '<b><a href="' + help_url + '" target="_blank">' + library_name + "</a></b>" 1628 if self.filename: 1629 dots = "" 1630 if len(self.filename) > 30: 1631 dots = "..." 1632 help_text += f"<br/><code><i>({dots}{self.filename[-30:]})</i></code>" 1633 1634 pdata = "" 1635 if self.dataset.GetPointData().GetScalars(): 1636 if self.dataset.GetPointData().GetScalars().GetName(): 1637 name = self.dataset.GetPointData().GetScalars().GetName() 1638 pdata = "<tr><td><b> point data array </b></td><td>" + name + "</td></tr>" 1639 1640 cdata = "" 1641 if self.dataset.GetCellData().GetScalars(): 1642 if self.dataset.GetCellData().GetScalars().GetName(): 1643 name = self.dataset.GetCellData().GetScalars().GetName() 1644 cdata = "<tr><td><b> cell data array </b></td><td>" + name + "</td></tr>" 1645 1646 pts = self.coordinates 1647 cm = np.mean(pts, axis=0) 1648 1649 _all = [ 1650 "<table>", 1651 "<tr>", 1652 "<td>", image, "</td>", 1653 "<td style='text-align: center; vertical-align: center;'><br/>", help_text, 1654 "<table>", 1655 "<tr><td><b> bounds </b> <br/> (x/y/z) </td><td>" + str(bounds) + "</td></tr>", 1656 "<tr><td><b> center of mass </b></td><td>" + utils.precision(cm,3) + "</td></tr>", 1657 "<tr><td><b> nr. points / cells </b></td><td>" 1658 + str(self.npoints) + " / " + str(self.ncells) + "</td></tr>", 1659 pdata, 1660 cdata, 1661 "</table>", 1662 "</table>", 1663 ] 1664 return "\n".join(_all) 1665 1666 def dimensions(self) -> np.ndarray: 1667 """Return the number of points in the x, y and z directions.""" 1668 return np.array(self.dataset.GetDimensions()) 1669 1670 def x_coordinates(self) -> np.ndarray: 1671 """Return the x-coordinates of the grid.""" 1672 return utils.vtk2numpy(self.dataset.GetXCoordinates()) 1673 1674 def y_coordinates(self) -> np.ndarray: 1675 """Return the y-coordinates of the grid.""" 1676 return utils.vtk2numpy(self.dataset.GetYCoordinates()) 1677 1678 def z_coordinates(self) -> np.ndarray: 1679 """Return the z-coordinates of the grid.""" 1680 return utils.vtk2numpy(self.dataset.GetZCoordinates()) 1681 1682 def is_point_visible(self, pid: int) -> bool: 1683 """Return True if point `pid` is visible.""" 1684 return self.dataset.IsPointVisible(pid) 1685 1686 def is_cell_visible(self, cid: int) -> bool: 1687 """Return True if cell `cid` is visible.""" 1688 return self.dataset.IsCellVisible(cid) 1689 1690 def has_blank_points(self) -> bool: 1691 """Return True if the grid has blank points.""" 1692 return self.dataset.HasAnyBlankPoints() 1693 1694 def has_blank_cells(self) -> bool: 1695 """Return True if the grid has blank cells.""" 1696 return self.dataset.HasAnyBlankCells() 1697 1698 def compute_structured_coords(self, x: list) -> dict: 1699 """ 1700 Convenience function computes the structured coordinates for a point `x`. 1701 1702 This method returns a dictionary with keys `ijk`, `pcoords` and `inside`. 1703 The cell is specified by the array `ijk`. 1704 and the parametric coordinates in the cell are specified with `pcoords`. 1705 Value of `inside` is False if the point x is outside of the grid. 1706 """ 1707 ijk = [0, 0, 0] 1708 pcoords = [0., 0., 0.] 1709 inout = self.dataset.ComputeStructuredCoordinates(x, ijk, pcoords) 1710 return {"ijk": np.array(ijk), "pcoords": np.array(pcoords), "inside": bool(inout)} 1711 1712 def compute_pointid(self, ijk: int) -> int: 1713 """Given a location in structured coordinates (i-j-k), return the point id.""" 1714 return self.dataset.ComputePointId(ijk) 1715 1716 def compute_cellid(self, ijk: int) -> int: 1717 """Given a location in structured coordinates (i-j-k), return the cell id.""" 1718 return self.dataset.ComputeCellId(ijk) 1719 1720 def find_point(self, x: list) -> int: 1721 """Given a position `x`, return the id of the closest point.""" 1722 return self.dataset.FindPoint(x) 1723 1724 def find_cell(self, x: list) -> dict: 1725 """Given a position `x`, return the id of the closest cell.""" 1726 cell = vtki.vtkHexagonalPrism() 1727 cellid = vtki.mutable(0) 1728 tol2 = 0.001 # vtki.mutable(0) 1729 subid = vtki.mutable(0) 1730 pcoords = [0.0, 0.0, 0.0] 1731 weights = [0.0, 0.0, 0.0] 1732 res = self.dataset.FindCell(x, cell, cellid, tol2, subid, pcoords, weights) 1733 result = {} 1734 result["cellid"] = cellid 1735 result["subid"] = subid 1736 result["pcoords"] = pcoords 1737 result["weights"] = weights 1738 result["status"] = res 1739 return result 1740 1741 def clone(self, deep=True) -> "RectilinearGrid": 1742 """Return a clone copy of the RectilinearGrid. Alias of `copy()`.""" 1743 if deep: 1744 newrg = vtki.vtkRectilinearGrid() 1745 newrg.CopyStructure(self.dataset) 1746 newrg.CopyAttributes(self.dataset) 1747 newvol = RectilinearGrid(newrg) 1748 else: 1749 newvol = RectilinearGrid(self.dataset) 1750 1751 prop = vtki.vtkProperty() 1752 prop.DeepCopy(self.properties) 1753 newvol.actor.SetProperty(prop) 1754 newvol.properties = prop 1755 newvol.pipeline = utils.OperationNode("clone", parents=[self], c="#bbd0ff", shape="diamond") 1756 return newvol 1757 1758 def bounds(self) -> np.ndarray: 1759 """ 1760 Get the object bounds. 1761 Returns a list in format `[xmin,xmax, ymin,ymax, zmin,zmax]`. 1762 """ 1763 # OVERRIDE CommonAlgorithms.bounds() which is too slow 1764 return np.array(self.dataset.GetBounds()) 1765 1766 def isosurface(self, value=None) -> "vedo.Mesh": 1767 """ 1768 Return a `Mesh` isosurface extracted from the object. 1769 1770 Set `value` as single float or list of values to draw the isosurface(s). 1771 """ 1772 scrange = self.dataset.GetScalarRange() 1773 1774 cf = vtki.new("ContourFilter") 1775 cf.UseScalarTreeOn() 1776 cf.SetInputData(self.dataset) 1777 cf.ComputeNormalsOn() 1778 1779 if value is None: 1780 value = (2 * scrange[0] + scrange[1]) / 3.0 1781 # print("automatic isosurface value =", value) 1782 cf.SetValue(0, value) 1783 else: 1784 if utils.is_sequence(value): 1785 cf.SetNumberOfContours(len(value)) 1786 for i, t in enumerate(value): 1787 cf.SetValue(i, t) 1788 else: 1789 cf.SetValue(0, value) 1790 1791 cf.Update() 1792 poly = cf.GetOutput() 1793 1794 out = vedo.mesh.Mesh(poly, c=None).phong() 1795 out.mapper.SetScalarRange(scrange[0], scrange[1]) 1796 1797 out.pipeline = utils.OperationNode( 1798 "isosurface", 1799 parents=[self], 1800 comment=f"#pts {out.dataset.GetNumberOfPoints()}", 1801 c="#4cc9f0:#e9c46a", 1802 ) 1803 return out 1804 1805 def cut_with_plane(self, origin=(0, 0, 0), normal="x") -> "UnstructuredGrid": 1806 """ 1807 Cut the object with the plane defined by a point and a normal. 1808 1809 Arguments: 1810 origin : (list) 1811 the cutting plane goes through this point 1812 normal : (list, str) 1813 normal vector to the cutting plane 1814 """ 1815 strn = str(normal) 1816 if strn == "x": normal = (1, 0, 0) 1817 elif strn == "y": normal = (0, 1, 0) 1818 elif strn == "z": normal = (0, 0, 1) 1819 elif strn == "-x": normal = (-1, 0, 0) 1820 elif strn == "-y": normal = (0, -1, 0) 1821 elif strn == "-z": normal = (0, 0, -1) 1822 plane = vtki.new("Plane") 1823 plane.SetOrigin(origin) 1824 plane.SetNormal(normal) 1825 clipper = vtki.new("ClipDataSet") 1826 clipper.SetInputData(self.dataset) 1827 clipper.SetClipFunction(plane) 1828 clipper.GenerateClipScalarsOff() 1829 clipper.GenerateClippedOutputOff() 1830 clipper.SetValue(0) 1831 clipper.Update() 1832 cout = clipper.GetOutput() 1833 ug = vedo.UnstructuredGrid(cout) 1834 if isinstance(self, UnstructuredGrid): 1835 self._update(cout) 1836 self.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 1837 return self 1838 ug.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 1839 return ug 1840 1841 def cut_with_mesh(self, mesh, invert=False, whole_cells=False, on_boundary=False) -> "UnstructuredGrid": 1842 """ 1843 Cut a `RectilinearGrid` with a `Mesh`. 1844 1845 Use `invert` to return cut off part of the input object. 1846 """ 1847 ug = self.dataset 1848 1849 ippd = vtki.new("ImplicitPolyDataDistance") 1850 ippd.SetInput(mesh.dataset) 1851 1852 if whole_cells or on_boundary: 1853 clipper = vtki.new("ExtractGeometry") 1854 clipper.SetInputData(ug) 1855 clipper.SetImplicitFunction(ippd) 1856 clipper.SetExtractInside(not invert) 1857 clipper.SetExtractBoundaryCells(False) 1858 if on_boundary: 1859 clipper.SetExtractBoundaryCells(True) 1860 clipper.SetExtractOnlyBoundaryCells(True) 1861 else: 1862 signed_dists = vtki.vtkFloatArray() 1863 signed_dists.SetNumberOfComponents(1) 1864 signed_dists.SetName("SignedDistance") 1865 for pointId in range(ug.GetNumberOfPoints()): 1866 p = ug.GetPoint(pointId) 1867 signed_dist = ippd.EvaluateFunction(p) 1868 signed_dists.InsertNextValue(signed_dist) 1869 ug.GetPointData().AddArray(signed_dists) 1870 ug.GetPointData().SetActiveScalars("SignedDistance") # NEEDED 1871 clipper = vtki.new("ClipDataSet") 1872 clipper.SetInputData(ug) 1873 clipper.SetInsideOut(not invert) 1874 clipper.SetValue(0.0) 1875 1876 clipper.Update() 1877 1878 out = UnstructuredGrid(clipper.GetOutput()) 1879 out.pipeline = utils.OperationNode("cut_with_mesh", parents=[self], c="#9e2a2b") 1880 return out
Build a rectilinear grid.
1405 def __init__(self, inputobj=None): 1406 """ 1407 A RectilinearGrid is a dataset where edges are parallel to the coordinate axes. 1408 It can be thought of as a tessellation of a box in 3D space, similar to a `Volume` 1409 except that the cells are not necessarily cubes, but they can have different lengths 1410 along each axis. 1411 This can be useful to describe a volume with variable resolution where one needs 1412 to represent a region with higher detail with respect to another region. 1413 1414 Arguments: 1415 inputobj : (vtkRectilinearGrid, list, str) 1416 list of points and indices, or filename 1417 1418 Example: 1419 ```python 1420 from vedo import RectilinearGrid, show 1421 1422 xcoords = 7 + np.sqrt(np.arange(0,2500,25)) 1423 ycoords = np.arange(0, 20) 1424 zcoords = np.arange(0, 20) 1425 1426 rgrid = RectilinearGrid([xcoords, ycoords, zcoords]) 1427 1428 print(rgrid) 1429 print(rgrid.x_coordinates().shape) 1430 print(rgrid.compute_structured_coords([20,10,11])) 1431 1432 msh = rgrid.tomesh().lw(1) 1433 1434 show(msh, axes=1, viewup="z") 1435 ``` 1436 """ 1437 1438 super().__init__() 1439 1440 self.dataset = None 1441 1442 self.mapper = vtki.new("PolyDataMapper") 1443 self._actor = vtki.vtkActor() 1444 self._actor.retrieve_object = weak_ref_to(self) 1445 self._actor.SetMapper(self.mapper) 1446 self.properties = self._actor.GetProperty() 1447 1448 self.transform = LinearTransform() 1449 self.point_locator = None 1450 self.cell_locator = None 1451 self.line_locator = None 1452 1453 self.name = "RectilinearGrid" 1454 self.filename = "" 1455 1456 self.info = {} 1457 self.time = time.time() 1458 1459 ############################### 1460 if inputobj is None: 1461 self.dataset = vtki.vtkRectilinearGrid() 1462 1463 elif isinstance(inputobj, vtki.vtkRectilinearGrid): 1464 self.dataset = inputobj 1465 1466 elif isinstance(inputobj, RectilinearGrid): 1467 self.dataset = inputobj.dataset 1468 1469 elif isinstance(inputobj, str): 1470 if "https://" in inputobj: 1471 inputobj = download(inputobj, verbose=False) 1472 if inputobj.endswith(".vtr"): 1473 reader = vtki.new("XMLRectilinearGridReader") 1474 else: 1475 reader = vtki.new("RectilinearGridReader") 1476 self.filename = inputobj 1477 reader.SetFileName(inputobj) 1478 reader.Update() 1479 self.dataset = reader.GetOutput() 1480 1481 elif utils.is_sequence(inputobj): 1482 self.dataset = vtki.vtkRectilinearGrid() 1483 xcoords, ycoords, zcoords = inputobj 1484 nx, ny, nz = len(xcoords), len(ycoords), len(zcoords) 1485 self.dataset.SetDimensions(nx, ny, nz) 1486 self.dataset.SetXCoordinates(utils.numpy2vtk(xcoords)) 1487 self.dataset.SetYCoordinates(utils.numpy2vtk(ycoords)) 1488 self.dataset.SetZCoordinates(utils.numpy2vtk(zcoords)) 1489 1490 ############################### 1491 1492 if not self.dataset: 1493 vedo.logger.error(f"RectilinearGrid: cannot understand input type {type(inputobj)}") 1494 return 1495 1496 self.properties.SetColor(0.352, 0.612, 0.996) # blue7 1497 1498 self.pipeline = utils.OperationNode( 1499 self, comment=f"#tets {self.dataset.GetNumberOfCells()}", c="#9e2a2b" 1500 )
A RectilinearGrid is a dataset where edges are parallel to the coordinate axes.
It can be thought of as a tessellation of a box in 3D space, similar to a Volume
except that the cells are not necessarily cubes, but they can have different lengths
along each axis.
This can be useful to describe a volume with variable resolution where one needs
to represent a region with higher detail with respect to another region.
Arguments:
- inputobj : (vtkRectilinearGrid, list, str) list of points and indices, or filename
Example:
from vedo import RectilinearGrid, show xcoords = 7 + np.sqrt(np.arange(0,2500,25)) ycoords = np.arange(0, 20) zcoords = np.arange(0, 20) rgrid = RectilinearGrid([xcoords, ycoords, zcoords]) print(rgrid) print(rgrid.x_coordinates().shape) print(rgrid.compute_structured_coords([20,10,11])) msh = rgrid.tomesh().lw(1) show(msh, axes=1, viewup="z")
1502 @property 1503 def actor(self): 1504 """Return the `vtkActor` of the object.""" 1505 gf = vtki.new("GeometryFilter") 1506 gf.SetInputData(self.dataset) 1507 gf.Update() 1508 self.mapper.SetInputData(gf.GetOutput()) 1509 self.mapper.Modified() 1510 return self._actor
Return the vtkActor of the object.
1666 def dimensions(self) -> np.ndarray: 1667 """Return the number of points in the x, y and z directions.""" 1668 return np.array(self.dataset.GetDimensions())
Return the number of points in the x, y and z directions.
1670 def x_coordinates(self) -> np.ndarray: 1671 """Return the x-coordinates of the grid.""" 1672 return utils.vtk2numpy(self.dataset.GetXCoordinates())
Return the x-coordinates of the grid.
1674 def y_coordinates(self) -> np.ndarray: 1675 """Return the y-coordinates of the grid.""" 1676 return utils.vtk2numpy(self.dataset.GetYCoordinates())
Return the y-coordinates of the grid.
1678 def z_coordinates(self) -> np.ndarray: 1679 """Return the z-coordinates of the grid.""" 1680 return utils.vtk2numpy(self.dataset.GetZCoordinates())
Return the z-coordinates of the grid.
1682 def is_point_visible(self, pid: int) -> bool: 1683 """Return True if point `pid` is visible.""" 1684 return self.dataset.IsPointVisible(pid)
Return True if point pid is visible.
1686 def is_cell_visible(self, cid: int) -> bool: 1687 """Return True if cell `cid` is visible.""" 1688 return self.dataset.IsCellVisible(cid)
Return True if cell cid is visible.
1690 def has_blank_points(self) -> bool: 1691 """Return True if the grid has blank points.""" 1692 return self.dataset.HasAnyBlankPoints()
Return True if the grid has blank points.
1694 def has_blank_cells(self) -> bool: 1695 """Return True if the grid has blank cells.""" 1696 return self.dataset.HasAnyBlankCells()
Return True if the grid has blank cells.
1698 def compute_structured_coords(self, x: list) -> dict: 1699 """ 1700 Convenience function computes the structured coordinates for a point `x`. 1701 1702 This method returns a dictionary with keys `ijk`, `pcoords` and `inside`. 1703 The cell is specified by the array `ijk`. 1704 and the parametric coordinates in the cell are specified with `pcoords`. 1705 Value of `inside` is False if the point x is outside of the grid. 1706 """ 1707 ijk = [0, 0, 0] 1708 pcoords = [0., 0., 0.] 1709 inout = self.dataset.ComputeStructuredCoordinates(x, ijk, pcoords) 1710 return {"ijk": np.array(ijk), "pcoords": np.array(pcoords), "inside": bool(inout)}
Convenience function computes the structured coordinates for a point x.
This method returns a dictionary with keys ijk, pcoords and inside.
The cell is specified by the array ijk.
and the parametric coordinates in the cell are specified with pcoords.
Value of inside is False if the point x is outside of the grid.
1712 def compute_pointid(self, ijk: int) -> int: 1713 """Given a location in structured coordinates (i-j-k), return the point id.""" 1714 return self.dataset.ComputePointId(ijk)
Given a location in structured coordinates (i-j-k), return the point id.
1716 def compute_cellid(self, ijk: int) -> int: 1717 """Given a location in structured coordinates (i-j-k), return the cell id.""" 1718 return self.dataset.ComputeCellId(ijk)
Given a location in structured coordinates (i-j-k), return the cell id.
1720 def find_point(self, x: list) -> int: 1721 """Given a position `x`, return the id of the closest point.""" 1722 return self.dataset.FindPoint(x)
Given a position x, return the id of the closest point.
1724 def find_cell(self, x: list) -> dict: 1725 """Given a position `x`, return the id of the closest cell.""" 1726 cell = vtki.vtkHexagonalPrism() 1727 cellid = vtki.mutable(0) 1728 tol2 = 0.001 # vtki.mutable(0) 1729 subid = vtki.mutable(0) 1730 pcoords = [0.0, 0.0, 0.0] 1731 weights = [0.0, 0.0, 0.0] 1732 res = self.dataset.FindCell(x, cell, cellid, tol2, subid, pcoords, weights) 1733 result = {} 1734 result["cellid"] = cellid 1735 result["subid"] = subid 1736 result["pcoords"] = pcoords 1737 result["weights"] = weights 1738 result["status"] = res 1739 return result
Given a position x, return the id of the closest cell.
1741 def clone(self, deep=True) -> "RectilinearGrid": 1742 """Return a clone copy of the RectilinearGrid. Alias of `copy()`.""" 1743 if deep: 1744 newrg = vtki.vtkRectilinearGrid() 1745 newrg.CopyStructure(self.dataset) 1746 newrg.CopyAttributes(self.dataset) 1747 newvol = RectilinearGrid(newrg) 1748 else: 1749 newvol = RectilinearGrid(self.dataset) 1750 1751 prop = vtki.vtkProperty() 1752 prop.DeepCopy(self.properties) 1753 newvol.actor.SetProperty(prop) 1754 newvol.properties = prop 1755 newvol.pipeline = utils.OperationNode("clone", parents=[self], c="#bbd0ff", shape="diamond") 1756 return newvol
Return a clone copy of the RectilinearGrid. Alias of copy().
1758 def bounds(self) -> np.ndarray: 1759 """ 1760 Get the object bounds. 1761 Returns a list in format `[xmin,xmax, ymin,ymax, zmin,zmax]`. 1762 """ 1763 # OVERRIDE CommonAlgorithms.bounds() which is too slow 1764 return np.array(self.dataset.GetBounds())
Get the object bounds.
Returns a list in format [xmin,xmax, ymin,ymax, zmin,zmax].
1766 def isosurface(self, value=None) -> "vedo.Mesh": 1767 """ 1768 Return a `Mesh` isosurface extracted from the object. 1769 1770 Set `value` as single float or list of values to draw the isosurface(s). 1771 """ 1772 scrange = self.dataset.GetScalarRange() 1773 1774 cf = vtki.new("ContourFilter") 1775 cf.UseScalarTreeOn() 1776 cf.SetInputData(self.dataset) 1777 cf.ComputeNormalsOn() 1778 1779 if value is None: 1780 value = (2 * scrange[0] + scrange[1]) / 3.0 1781 # print("automatic isosurface value =", value) 1782 cf.SetValue(0, value) 1783 else: 1784 if utils.is_sequence(value): 1785 cf.SetNumberOfContours(len(value)) 1786 for i, t in enumerate(value): 1787 cf.SetValue(i, t) 1788 else: 1789 cf.SetValue(0, value) 1790 1791 cf.Update() 1792 poly = cf.GetOutput() 1793 1794 out = vedo.mesh.Mesh(poly, c=None).phong() 1795 out.mapper.SetScalarRange(scrange[0], scrange[1]) 1796 1797 out.pipeline = utils.OperationNode( 1798 "isosurface", 1799 parents=[self], 1800 comment=f"#pts {out.dataset.GetNumberOfPoints()}", 1801 c="#4cc9f0:#e9c46a", 1802 ) 1803 return out
Return a Mesh isosurface extracted from the object.
Set value as single float or list of values to draw the isosurface(s).
1805 def cut_with_plane(self, origin=(0, 0, 0), normal="x") -> "UnstructuredGrid": 1806 """ 1807 Cut the object with the plane defined by a point and a normal. 1808 1809 Arguments: 1810 origin : (list) 1811 the cutting plane goes through this point 1812 normal : (list, str) 1813 normal vector to the cutting plane 1814 """ 1815 strn = str(normal) 1816 if strn == "x": normal = (1, 0, 0) 1817 elif strn == "y": normal = (0, 1, 0) 1818 elif strn == "z": normal = (0, 0, 1) 1819 elif strn == "-x": normal = (-1, 0, 0) 1820 elif strn == "-y": normal = (0, -1, 0) 1821 elif strn == "-z": normal = (0, 0, -1) 1822 plane = vtki.new("Plane") 1823 plane.SetOrigin(origin) 1824 plane.SetNormal(normal) 1825 clipper = vtki.new("ClipDataSet") 1826 clipper.SetInputData(self.dataset) 1827 clipper.SetClipFunction(plane) 1828 clipper.GenerateClipScalarsOff() 1829 clipper.GenerateClippedOutputOff() 1830 clipper.SetValue(0) 1831 clipper.Update() 1832 cout = clipper.GetOutput() 1833 ug = vedo.UnstructuredGrid(cout) 1834 if isinstance(self, UnstructuredGrid): 1835 self._update(cout) 1836 self.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 1837 return self 1838 ug.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 1839 return ug
Cut the object with the plane defined by a point and a normal.
Arguments:
- origin : (list) the cutting plane goes through this point
- normal : (list, str) normal vector to the cutting plane
1841 def cut_with_mesh(self, mesh, invert=False, whole_cells=False, on_boundary=False) -> "UnstructuredGrid": 1842 """ 1843 Cut a `RectilinearGrid` with a `Mesh`. 1844 1845 Use `invert` to return cut off part of the input object. 1846 """ 1847 ug = self.dataset 1848 1849 ippd = vtki.new("ImplicitPolyDataDistance") 1850 ippd.SetInput(mesh.dataset) 1851 1852 if whole_cells or on_boundary: 1853 clipper = vtki.new("ExtractGeometry") 1854 clipper.SetInputData(ug) 1855 clipper.SetImplicitFunction(ippd) 1856 clipper.SetExtractInside(not invert) 1857 clipper.SetExtractBoundaryCells(False) 1858 if on_boundary: 1859 clipper.SetExtractBoundaryCells(True) 1860 clipper.SetExtractOnlyBoundaryCells(True) 1861 else: 1862 signed_dists = vtki.vtkFloatArray() 1863 signed_dists.SetNumberOfComponents(1) 1864 signed_dists.SetName("SignedDistance") 1865 for pointId in range(ug.GetNumberOfPoints()): 1866 p = ug.GetPoint(pointId) 1867 signed_dist = ippd.EvaluateFunction(p) 1868 signed_dists.InsertNextValue(signed_dist) 1869 ug.GetPointData().AddArray(signed_dists) 1870 ug.GetPointData().SetActiveScalars("SignedDistance") # NEEDED 1871 clipper = vtki.new("ClipDataSet") 1872 clipper.SetInputData(ug) 1873 clipper.SetInsideOut(not invert) 1874 clipper.SetValue(0.0) 1875 1876 clipper.Update() 1877 1878 out = UnstructuredGrid(clipper.GetOutput()) 1879 out.pipeline = utils.OperationNode("cut_with_mesh", parents=[self], c="#9e2a2b") 1880 return out
Cut a RectilinearGrid with a Mesh.
Use invert to return cut off part of the input object.
Inherited Members
- vedo.core.PointAlgorithms
- apply_transform
- apply_transform_from_actor
- pos
- shift
- x
- y
- z
- rotate
- rotate_x
- rotate_y
- rotate_z
- reorient
- scale
- vedo.core.CommonAlgorithms
- pointdata
- celldata
- metadata
- rename
- memory_address
- memory_size
- modified
- box
- update_dataset
- xbounds
- ybounds
- zbounds
- diagonal_size
- average_size
- center_of_mass
- copy_data_from
- inputdata
- npoints
- nvertices
- ncells
- cell_centers
- lines
- lines_as_flat_array
- mark_boundaries
- find_cells_in_bounds
- find_cells_along_line
- find_cells_along_plane
- keep_cell_types
- map_cells_to_points
- vertices
- points
- coordinates
- cells_as_flat_array
- cells
- cell_edge_neighbors
- map_points_to_cells
- resample_data_from
- interpolate_data_from
- add_ids
- gradient
- divergence
- vorticity
- probe
- compute_cell_size
- generate_random_data
- integrate_data
- write
- tomesh
- signed_distance
- unsigned_distance
- smooth_data
- compute_streamlines
- vedo.visual.MeshVisual
- follow_camera
- wireframe
- flat
- phong
- backface_culling
- render_lines_as_tubes
- frontface_culling
- backcolor
- bc
- linewidth
- lw
- linecolor
- lc
- texture
- vedo.visual.PointsVisual
- clone2d
- copy_properties_from
- color
- c
- alpha
- lut_color_at
- opacity
- force_opaque
- force_translucent
- point_size
- ps
- render_points_as_spheres
- lighting
- point_blurring
- cellcolors
- pointcolors
- cmap
- add_trail
- update_trail
- add_shadow
- update_shadows
- labels
- labels2d
- legend
- flagpole
- flagpost
- caption
1884class StructuredGrid(PointAlgorithms, MeshVisual): 1885 """ 1886 Build a structured grid. 1887 """ 1888 1889 def __init__(self, inputobj=None): 1890 """ 1891 A StructuredGrid is a dataset where edges of the hexahedrons are 1892 not necessarily parallel to the coordinate axes. 1893 It can be thought of as a tessellation of a block of 3D space, 1894 similar to a `RectilinearGrid` 1895 except that the cells are not necessarily cubes, they can have different 1896 orientations but are connected in the same way as a `RectilinearGrid`. 1897 1898 Arguments: 1899 inputobj : (vtkStructuredGrid, list, str) 1900 list of points and indices, or filename 1901 1902 Example: 1903 ```python 1904 from vedo import * 1905 sgrid = StructuredGrid(dataurl+"structgrid.vts") 1906 print(sgrid) 1907 msh = sgrid.tomesh().lw(1) 1908 show(msh, axes=1, viewup="z") 1909 ``` 1910 1911 ```python 1912 from vedo import * 1913 1914 cx = np.sqrt(np.linspace(100, 400, 10)) 1915 cy = np.linspace(30, 40, 20) 1916 cz = np.linspace(40, 50, 30) 1917 x, y, z = np.meshgrid(cx, cy, cz) 1918 1919 sgrid1 = StructuredGrid([x, y, z]) 1920 sgrid1.cmap("viridis", sgrid1.coordinates[:, 0]) 1921 print(sgrid1) 1922 1923 sgrid2 = sgrid1.clone().cut_with_plane(normal=(-1,1,1), origin=[14,34,44]) 1924 msh2 = sgrid2.tomesh(shrink=0.9).lw(1).cmap("viridis") 1925 1926 show( 1927 [["StructuredGrid", sgrid1], ["Shrinked Mesh", msh2]], 1928 N=2, axes=1, viewup="z", 1929 ) 1930 ``` 1931 """ 1932 1933 super().__init__() 1934 1935 self.dataset = None 1936 1937 self.mapper = vtki.new("PolyDataMapper") 1938 self._actor = vtki.vtkActor() 1939 self._actor.retrieve_object = weak_ref_to(self) 1940 self._actor.SetMapper(self.mapper) 1941 self.properties = self._actor.GetProperty() 1942 1943 self.transform = LinearTransform() 1944 self.point_locator = None 1945 self.cell_locator = None 1946 self.line_locator = None 1947 1948 self.name = "StructuredGrid" 1949 self.filename = "" 1950 1951 self.info = {} 1952 self.time = time.time() 1953 1954 ############################### 1955 if inputobj is None: 1956 self.dataset = vtki.vtkStructuredGrid() 1957 1958 elif isinstance(inputobj, vtki.vtkStructuredGrid): 1959 self.dataset = inputobj 1960 1961 elif isinstance(inputobj, StructuredGrid): 1962 self.dataset = inputobj.dataset 1963 1964 elif isinstance(inputobj, str): 1965 if "https://" in inputobj: 1966 inputobj = download(inputobj, verbose=False) 1967 if inputobj.endswith(".vts"): 1968 reader = vtki.new("XMLStructuredGridReader") 1969 else: 1970 reader = vtki.new("StructuredGridReader") 1971 self.filename = inputobj 1972 reader.SetFileName(inputobj) 1973 reader.Update() 1974 self.dataset = reader.GetOutput() 1975 1976 elif utils.is_sequence(inputobj): 1977 self.dataset = vtki.vtkStructuredGrid() 1978 x, y, z = inputobj 1979 xyz = np.vstack(( 1980 x.flatten(order="F"), 1981 y.flatten(order="F"), 1982 z.flatten(order="F")) 1983 ).T 1984 dims = x.shape 1985 self.dataset.SetDimensions(dims) 1986 # self.dataset.SetDimensions(dims[1], dims[0], dims[2]) 1987 vpoints = vtki.vtkPoints() 1988 vpoints.SetData(utils.numpy2vtk(xyz)) 1989 self.dataset.SetPoints(vpoints) 1990 1991 1992 ############################### 1993 if not self.dataset: 1994 vedo.logger.error(f"StructuredGrid: cannot understand input type {type(inputobj)}") 1995 return 1996 1997 self.properties.SetColor(0.352, 0.612, 0.996) # blue7 1998 1999 self.pipeline = utils.OperationNode( 2000 self, comment=f"#tets {self.dataset.GetNumberOfCells()}", c="#9e2a2b" 2001 ) 2002 2003 @property 2004 def actor(self): 2005 """Return the `vtkActor` of the object.""" 2006 gf = vtki.new("GeometryFilter") 2007 gf.SetInputData(self.dataset) 2008 gf.Update() 2009 self.mapper.SetInputData(gf.GetOutput()) 2010 self.mapper.Modified() 2011 return self._actor 2012 2013 @actor.setter 2014 def actor(self, _): 2015 pass 2016 2017 def _update(self, data, reset_locators=False): 2018 self.dataset = data 2019 if reset_locators: 2020 self.cell_locator = None 2021 self.point_locator = None 2022 return self 2023 2024 ################################################################## 2025 def __str__(self): 2026 """Print a summary for the `StructuredGrid` object.""" 2027 module = self.__class__.__module__ 2028 name = self.__class__.__name__ 2029 out = vedo.printc( 2030 f"{module}.{name} at ({hex(self.memory_address())})".ljust(75), 2031 c="c", bold=True, invert=True, return_string=True, 2032 ) 2033 out += "\x1b[0m\x1b[36;1m" 2034 2035 out += "name".ljust(14) + ": " + str(self.name) + "\n" 2036 if self.filename: 2037 out += "filename".ljust(14) + ": " + str(self.filename) + "\n" 2038 2039 out += "dimensions".ljust(14) + ": " + str(self.dataset.GetDimensions()) + "\n" 2040 2041 out += "center".ljust(14) + ": " 2042 out += utils.precision(self.dataset.GetCenter(), 6) + "\n" 2043 2044 bnds = self.bounds() 2045 bx1, bx2 = utils.precision(bnds[0], 3), utils.precision(bnds[1], 3) 2046 by1, by2 = utils.precision(bnds[2], 3), utils.precision(bnds[3], 3) 2047 bz1, bz2 = utils.precision(bnds[4], 3), utils.precision(bnds[5], 3) 2048 out += "bounds".ljust(14) + ":" 2049 out += " x=(" + bx1 + ", " + bx2 + ")," 2050 out += " y=(" + by1 + ", " + by2 + ")," 2051 out += " z=(" + bz1 + ", " + bz2 + ")\n" 2052 2053 out += "memory size".ljust(14) + ": " 2054 out += utils.precision(self.dataset.GetActualMemorySize() / 1024, 2) + " MB\n" 2055 2056 for key in self.pointdata.keys(): 2057 arr = self.pointdata[key] 2058 rng = utils.precision(arr.min(), 3) + ", " + utils.precision(arr.max(), 3) 2059 mark_active = "pointdata" 2060 a_scalars = self.dataset.GetPointData().GetScalars() 2061 a_vectors = self.dataset.GetPointData().GetVectors() 2062 a_tensors = self.dataset.GetPointData().GetTensors() 2063 if a_scalars and a_scalars.GetName() == key: 2064 mark_active += " *" 2065 elif a_vectors and a_vectors.GetName() == key: 2066 mark_active += " **" 2067 elif a_tensors and a_tensors.GetName() == key: 2068 mark_active += " ***" 2069 out += mark_active.ljust(14) + f': "{key}" ({arr.dtype}), ndim={arr.ndim}' 2070 out += f", range=({rng})\n" 2071 2072 for key in self.celldata.keys(): 2073 arr = self.celldata[key] 2074 rng = utils.precision(arr.min(), 3) + ", " + utils.precision(arr.max(), 3) 2075 mark_active = "celldata" 2076 a_scalars = self.dataset.GetCellData().GetScalars() 2077 a_vectors = self.dataset.GetCellData().GetVectors() 2078 a_tensors = self.dataset.GetCellData().GetTensors() 2079 if a_scalars and a_scalars.GetName() == key: 2080 mark_active += " *" 2081 elif a_vectors and a_vectors.GetName() == key: 2082 mark_active += " **" 2083 elif a_tensors and a_tensors.GetName() == key: 2084 mark_active += " ***" 2085 out += mark_active.ljust(14) + f': "{key}" ({arr.dtype}), ndim={arr.ndim}' 2086 out += f", range=({rng})\n" 2087 2088 for key in self.metadata.keys(): 2089 arr = self.metadata[key] 2090 out += "metadata".ljust(14) + ": " + f'"{key}" ({len(arr)} values)\n' 2091 2092 return out.rstrip() + "\x1b[0m" 2093 2094 def _repr_html_(self): 2095 """ 2096 HTML representation of the StructuredGrid object for Jupyter Notebooks. 2097 2098 Returns: 2099 HTML text with the image and some properties. 2100 """ 2101 import io 2102 import base64 2103 from PIL import Image 2104 2105 library_name = "vedo.grids.StructuredGrid" 2106 help_url = "https://vedo.embl.es/docs/vedo/grids.html#StructuredGrid" 2107 2108 m = self.tomesh().linewidth(1).lighting("off") 2109 arr= m.thumbnail(zoom=1, elevation=-30, azimuth=-30) 2110 2111 im = Image.fromarray(arr) 2112 buffered = io.BytesIO() 2113 im.save(buffered, format="PNG", quality=100) 2114 encoded = base64.b64encode(buffered.getvalue()).decode("utf-8") 2115 url = "data:image/png;base64," + encoded 2116 image = f"<img src='{url}'></img>" 2117 2118 bounds = "<br/>".join( 2119 [ 2120 utils.precision(min_x, 4) + " ... " + utils.precision(max_x, 4) 2121 for min_x, max_x in zip(self.bounds()[::2], self.bounds()[1::2]) 2122 ] 2123 ) 2124 2125 help_text = "" 2126 if self.name: 2127 help_text += f"<b> {self.name}:   </b>" 2128 help_text += '<b><a href="' + help_url + '" target="_blank">' + library_name + "</a></b>" 2129 if self.filename: 2130 dots = "" 2131 if len(self.filename) > 30: 2132 dots = "..." 2133 help_text += f"<br/><code><i>({dots}{self.filename[-30:]})</i></code>" 2134 2135 pdata = "" 2136 if self.dataset.GetPointData().GetScalars(): 2137 if self.dataset.GetPointData().GetScalars().GetName(): 2138 name = self.dataset.GetPointData().GetScalars().GetName() 2139 pdata = "<tr><td><b> point data array </b></td><td>" + name + "</td></tr>" 2140 2141 cdata = "" 2142 if self.dataset.GetCellData().GetScalars(): 2143 if self.dataset.GetCellData().GetScalars().GetName(): 2144 name = self.dataset.GetCellData().GetScalars().GetName() 2145 cdata = "<tr><td><b> cell data array </b></td><td>" + name + "</td></tr>" 2146 2147 pts = self.coordinates 2148 cm = np.mean(pts, axis=0) 2149 2150 _all = [ 2151 "<table>", 2152 "<tr>", 2153 "<td>", image, "</td>", 2154 "<td style='text-align: center; vertical-align: center;'><br/>", help_text, 2155 "<table>", 2156 "<tr><td><b> bounds </b> <br/> (x/y/z) </td><td>" + str(bounds) + "</td></tr>", 2157 "<tr><td><b> center of mass </b></td><td>" + utils.precision(cm,3) + "</td></tr>", 2158 "<tr><td><b> nr. points / cells </b></td><td>" 2159 + str(self.npoints) + " / " + str(self.ncells) + "</td></tr>", 2160 pdata, 2161 cdata, 2162 "</table>", 2163 "</table>", 2164 ] 2165 return "\n".join(_all) 2166 2167 def dimensions(self) -> np.ndarray: 2168 """Return the number of points in the x, y and z directions.""" 2169 return np.array(self.dataset.GetDimensions()) 2170 2171 def clone(self, deep=True) -> "StructuredGrid": 2172 """Return a clone copy of the StructuredGrid. Alias of `copy()`.""" 2173 if deep: 2174 newrg = vtki.vtkStructuredGrid() 2175 newrg.CopyStructure(self.dataset) 2176 newrg.CopyAttributes(self.dataset) 2177 newvol = StructuredGrid(newrg) 2178 else: 2179 newvol = StructuredGrid(self.dataset) 2180 2181 prop = vtki.vtkProperty() 2182 prop.DeepCopy(self.properties) 2183 newvol.actor.SetProperty(prop) 2184 newvol.properties = prop 2185 newvol.pipeline = utils.OperationNode("clone", parents=[self], c="#bbd0ff", shape="diamond") 2186 return newvol 2187 2188 def find_point(self, x: list) -> int: 2189 """Given a position `x`, return the id of the closest point.""" 2190 return self.dataset.FindPoint(x) 2191 2192 def find_cell(self, x: list) -> dict: 2193 """Given a position `x`, return the id of the closest cell.""" 2194 cell = vtki.vtkHexagonalPrism() 2195 cellid = vtki.mutable(0) 2196 tol2 = 0.001 # vtki.mutable(0) 2197 subid = vtki.mutable(0) 2198 pcoords = [0.0, 0.0, 0.0] 2199 weights = [0.0, 0.0, 0.0] 2200 res = self.dataset.FindCell(x, cell, cellid, tol2, subid, pcoords, weights) 2201 result = {} 2202 result["cellid"] = cellid 2203 result["subid"] = subid 2204 result["pcoords"] = pcoords 2205 result["weights"] = weights 2206 result["status"] = res 2207 return result 2208 2209 def cut_with_plane(self, origin=(0, 0, 0), normal="x") -> "vedo.UnstructuredGrid": 2210 """ 2211 Cut the object with the plane defined by a point and a normal. 2212 2213 Arguments: 2214 origin : (list) 2215 the cutting plane goes through this point 2216 normal : (list, str) 2217 normal vector to the cutting plane 2218 2219 Returns an `UnstructuredGrid` object. 2220 """ 2221 strn = str(normal) 2222 if strn == "x": normal = (1, 0, 0) 2223 elif strn == "y": normal = (0, 1, 0) 2224 elif strn == "z": normal = (0, 0, 1) 2225 elif strn == "-x": normal = (-1, 0, 0) 2226 elif strn == "-y": normal = (0, -1, 0) 2227 elif strn == "-z": normal = (0, 0, -1) 2228 plane = vtki.new("Plane") 2229 plane.SetOrigin(origin) 2230 plane.SetNormal(normal) 2231 clipper = vtki.new("ClipDataSet") 2232 clipper.SetInputData(self.dataset) 2233 clipper.SetClipFunction(plane) 2234 clipper.GenerateClipScalarsOff() 2235 clipper.GenerateClippedOutputOff() 2236 clipper.SetValue(0) 2237 clipper.Update() 2238 cout = clipper.GetOutput() 2239 ug = vedo.UnstructuredGrid(cout) 2240 if isinstance(self, vedo.UnstructuredGrid): 2241 self._update(cout) 2242 self.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 2243 return self 2244 ug.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 2245 return ug 2246 2247 def cut_with_mesh(self, mesh: Mesh, invert=False, whole_cells=False, on_boundary=False) -> "UnstructuredGrid": 2248 """ 2249 Cut a `RectilinearGrid` with a `Mesh`. 2250 2251 Use `invert` to return cut off part of the input object. 2252 2253 Returns an `UnstructuredGrid` object. 2254 """ 2255 ug = self.dataset 2256 2257 ippd = vtki.new("ImplicitPolyDataDistance") 2258 ippd.SetInput(mesh.dataset) 2259 2260 if whole_cells or on_boundary: 2261 clipper = vtki.new("ExtractGeometry") 2262 clipper.SetInputData(ug) 2263 clipper.SetImplicitFunction(ippd) 2264 clipper.SetExtractInside(not invert) 2265 clipper.SetExtractBoundaryCells(False) 2266 if on_boundary: 2267 clipper.SetExtractBoundaryCells(True) 2268 clipper.SetExtractOnlyBoundaryCells(True) 2269 else: 2270 signed_dists = vtki.vtkFloatArray() 2271 signed_dists.SetNumberOfComponents(1) 2272 signed_dists.SetName("SignedDistance") 2273 for pointId in range(ug.GetNumberOfPoints()): 2274 p = ug.GetPoint(pointId) 2275 signed_dist = ippd.EvaluateFunction(p) 2276 signed_dists.InsertNextValue(signed_dist) 2277 ug.GetPointData().AddArray(signed_dists) 2278 ug.GetPointData().SetActiveScalars("SignedDistance") # NEEDED 2279 clipper = vtki.new("ClipDataSet") 2280 clipper.SetInputData(ug) 2281 clipper.SetInsideOut(not invert) 2282 clipper.SetValue(0.0) 2283 2284 clipper.Update() 2285 2286 out = UnstructuredGrid(clipper.GetOutput()) 2287 out.pipeline = utils.OperationNode("cut_with_mesh", parents=[self], c="#9e2a2b") 2288 return out 2289 2290 def isosurface(self, value=None) -> "vedo.Mesh": 2291 """ 2292 Return a `Mesh` isosurface extracted from the object. 2293 2294 Set `value` as single float or list of values to draw the isosurface(s). 2295 """ 2296 scrange = self.dataset.GetScalarRange() 2297 2298 cf = vtki.new("ContourFilter") 2299 cf.UseScalarTreeOn() 2300 cf.SetInputData(self.dataset) 2301 cf.ComputeNormalsOn() 2302 2303 if value is None: 2304 value = (2 * scrange[0] + scrange[1]) / 3.0 2305 # print("automatic isosurface value =", value) 2306 cf.SetValue(0, value) 2307 else: 2308 if utils.is_sequence(value): 2309 cf.SetNumberOfContours(len(value)) 2310 for i, t in enumerate(value): 2311 cf.SetValue(i, t) 2312 else: 2313 cf.SetValue(0, value) 2314 2315 cf.Update() 2316 poly = cf.GetOutput() 2317 2318 out = vedo.mesh.Mesh(poly, c=None).phong() 2319 out.mapper.SetScalarRange(scrange[0], scrange[1]) 2320 2321 out.pipeline = utils.OperationNode( 2322 "isosurface", 2323 parents=[self], 2324 comment=f"#pts {out.dataset.GetNumberOfPoints()}", 2325 c="#4cc9f0:#e9c46a", 2326 ) 2327 return out
Build a structured grid.
1889 def __init__(self, inputobj=None): 1890 """ 1891 A StructuredGrid is a dataset where edges of the hexahedrons are 1892 not necessarily parallel to the coordinate axes. 1893 It can be thought of as a tessellation of a block of 3D space, 1894 similar to a `RectilinearGrid` 1895 except that the cells are not necessarily cubes, they can have different 1896 orientations but are connected in the same way as a `RectilinearGrid`. 1897 1898 Arguments: 1899 inputobj : (vtkStructuredGrid, list, str) 1900 list of points and indices, or filename 1901 1902 Example: 1903 ```python 1904 from vedo import * 1905 sgrid = StructuredGrid(dataurl+"structgrid.vts") 1906 print(sgrid) 1907 msh = sgrid.tomesh().lw(1) 1908 show(msh, axes=1, viewup="z") 1909 ``` 1910 1911 ```python 1912 from vedo import * 1913 1914 cx = np.sqrt(np.linspace(100, 400, 10)) 1915 cy = np.linspace(30, 40, 20) 1916 cz = np.linspace(40, 50, 30) 1917 x, y, z = np.meshgrid(cx, cy, cz) 1918 1919 sgrid1 = StructuredGrid([x, y, z]) 1920 sgrid1.cmap("viridis", sgrid1.coordinates[:, 0]) 1921 print(sgrid1) 1922 1923 sgrid2 = sgrid1.clone().cut_with_plane(normal=(-1,1,1), origin=[14,34,44]) 1924 msh2 = sgrid2.tomesh(shrink=0.9).lw(1).cmap("viridis") 1925 1926 show( 1927 [["StructuredGrid", sgrid1], ["Shrinked Mesh", msh2]], 1928 N=2, axes=1, viewup="z", 1929 ) 1930 ``` 1931 """ 1932 1933 super().__init__() 1934 1935 self.dataset = None 1936 1937 self.mapper = vtki.new("PolyDataMapper") 1938 self._actor = vtki.vtkActor() 1939 self._actor.retrieve_object = weak_ref_to(self) 1940 self._actor.SetMapper(self.mapper) 1941 self.properties = self._actor.GetProperty() 1942 1943 self.transform = LinearTransform() 1944 self.point_locator = None 1945 self.cell_locator = None 1946 self.line_locator = None 1947 1948 self.name = "StructuredGrid" 1949 self.filename = "" 1950 1951 self.info = {} 1952 self.time = time.time() 1953 1954 ############################### 1955 if inputobj is None: 1956 self.dataset = vtki.vtkStructuredGrid() 1957 1958 elif isinstance(inputobj, vtki.vtkStructuredGrid): 1959 self.dataset = inputobj 1960 1961 elif isinstance(inputobj, StructuredGrid): 1962 self.dataset = inputobj.dataset 1963 1964 elif isinstance(inputobj, str): 1965 if "https://" in inputobj: 1966 inputobj = download(inputobj, verbose=False) 1967 if inputobj.endswith(".vts"): 1968 reader = vtki.new("XMLStructuredGridReader") 1969 else: 1970 reader = vtki.new("StructuredGridReader") 1971 self.filename = inputobj 1972 reader.SetFileName(inputobj) 1973 reader.Update() 1974 self.dataset = reader.GetOutput() 1975 1976 elif utils.is_sequence(inputobj): 1977 self.dataset = vtki.vtkStructuredGrid() 1978 x, y, z = inputobj 1979 xyz = np.vstack(( 1980 x.flatten(order="F"), 1981 y.flatten(order="F"), 1982 z.flatten(order="F")) 1983 ).T 1984 dims = x.shape 1985 self.dataset.SetDimensions(dims) 1986 # self.dataset.SetDimensions(dims[1], dims[0], dims[2]) 1987 vpoints = vtki.vtkPoints() 1988 vpoints.SetData(utils.numpy2vtk(xyz)) 1989 self.dataset.SetPoints(vpoints) 1990 1991 1992 ############################### 1993 if not self.dataset: 1994 vedo.logger.error(f"StructuredGrid: cannot understand input type {type(inputobj)}") 1995 return 1996 1997 self.properties.SetColor(0.352, 0.612, 0.996) # blue7 1998 1999 self.pipeline = utils.OperationNode( 2000 self, comment=f"#tets {self.dataset.GetNumberOfCells()}", c="#9e2a2b" 2001 )
A StructuredGrid is a dataset where edges of the hexahedrons are
not necessarily parallel to the coordinate axes.
It can be thought of as a tessellation of a block of 3D space,
similar to a RectilinearGrid
except that the cells are not necessarily cubes, they can have different
orientations but are connected in the same way as a RectilinearGrid.
Arguments:
- inputobj : (vtkStructuredGrid, list, str) list of points and indices, or filename
Example:
from vedo import * sgrid = StructuredGrid(dataurl+"structgrid.vts") print(sgrid) msh = sgrid.tomesh().lw(1) show(msh, axes=1, viewup="z")from vedo import * cx = np.sqrt(np.linspace(100, 400, 10)) cy = np.linspace(30, 40, 20) cz = np.linspace(40, 50, 30) x, y, z = np.meshgrid(cx, cy, cz) sgrid1 = StructuredGrid([x, y, z]) sgrid1.cmap("viridis", sgrid1.coordinates[:, 0]) print(sgrid1) sgrid2 = sgrid1.clone().cut_with_plane(normal=(-1,1,1), origin=[14,34,44]) msh2 = sgrid2.tomesh(shrink=0.9).lw(1).cmap("viridis") show( [["StructuredGrid", sgrid1], ["Shrinked Mesh", msh2]], N=2, axes=1, viewup="z", )
2003 @property 2004 def actor(self): 2005 """Return the `vtkActor` of the object.""" 2006 gf = vtki.new("GeometryFilter") 2007 gf.SetInputData(self.dataset) 2008 gf.Update() 2009 self.mapper.SetInputData(gf.GetOutput()) 2010 self.mapper.Modified() 2011 return self._actor
Return the vtkActor of the object.
2167 def dimensions(self) -> np.ndarray: 2168 """Return the number of points in the x, y and z directions.""" 2169 return np.array(self.dataset.GetDimensions())
Return the number of points in the x, y and z directions.
2171 def clone(self, deep=True) -> "StructuredGrid": 2172 """Return a clone copy of the StructuredGrid. Alias of `copy()`.""" 2173 if deep: 2174 newrg = vtki.vtkStructuredGrid() 2175 newrg.CopyStructure(self.dataset) 2176 newrg.CopyAttributes(self.dataset) 2177 newvol = StructuredGrid(newrg) 2178 else: 2179 newvol = StructuredGrid(self.dataset) 2180 2181 prop = vtki.vtkProperty() 2182 prop.DeepCopy(self.properties) 2183 newvol.actor.SetProperty(prop) 2184 newvol.properties = prop 2185 newvol.pipeline = utils.OperationNode("clone", parents=[self], c="#bbd0ff", shape="diamond") 2186 return newvol
Return a clone copy of the StructuredGrid. Alias of copy().
2188 def find_point(self, x: list) -> int: 2189 """Given a position `x`, return the id of the closest point.""" 2190 return self.dataset.FindPoint(x)
Given a position x, return the id of the closest point.
2192 def find_cell(self, x: list) -> dict: 2193 """Given a position `x`, return the id of the closest cell.""" 2194 cell = vtki.vtkHexagonalPrism() 2195 cellid = vtki.mutable(0) 2196 tol2 = 0.001 # vtki.mutable(0) 2197 subid = vtki.mutable(0) 2198 pcoords = [0.0, 0.0, 0.0] 2199 weights = [0.0, 0.0, 0.0] 2200 res = self.dataset.FindCell(x, cell, cellid, tol2, subid, pcoords, weights) 2201 result = {} 2202 result["cellid"] = cellid 2203 result["subid"] = subid 2204 result["pcoords"] = pcoords 2205 result["weights"] = weights 2206 result["status"] = res 2207 return result
Given a position x, return the id of the closest cell.
2209 def cut_with_plane(self, origin=(0, 0, 0), normal="x") -> "vedo.UnstructuredGrid": 2210 """ 2211 Cut the object with the plane defined by a point and a normal. 2212 2213 Arguments: 2214 origin : (list) 2215 the cutting plane goes through this point 2216 normal : (list, str) 2217 normal vector to the cutting plane 2218 2219 Returns an `UnstructuredGrid` object. 2220 """ 2221 strn = str(normal) 2222 if strn == "x": normal = (1, 0, 0) 2223 elif strn == "y": normal = (0, 1, 0) 2224 elif strn == "z": normal = (0, 0, 1) 2225 elif strn == "-x": normal = (-1, 0, 0) 2226 elif strn == "-y": normal = (0, -1, 0) 2227 elif strn == "-z": normal = (0, 0, -1) 2228 plane = vtki.new("Plane") 2229 plane.SetOrigin(origin) 2230 plane.SetNormal(normal) 2231 clipper = vtki.new("ClipDataSet") 2232 clipper.SetInputData(self.dataset) 2233 clipper.SetClipFunction(plane) 2234 clipper.GenerateClipScalarsOff() 2235 clipper.GenerateClippedOutputOff() 2236 clipper.SetValue(0) 2237 clipper.Update() 2238 cout = clipper.GetOutput() 2239 ug = vedo.UnstructuredGrid(cout) 2240 if isinstance(self, vedo.UnstructuredGrid): 2241 self._update(cout) 2242 self.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 2243 return self 2244 ug.pipeline = utils.OperationNode("cut_with_plane", parents=[self], c="#9e2a2b") 2245 return ug
Cut the object with the plane defined by a point and a normal.
Arguments:
- origin : (list) the cutting plane goes through this point
- normal : (list, str) normal vector to the cutting plane
Returns an UnstructuredGrid object.
2247 def cut_with_mesh(self, mesh: Mesh, invert=False, whole_cells=False, on_boundary=False) -> "UnstructuredGrid": 2248 """ 2249 Cut a `RectilinearGrid` with a `Mesh`. 2250 2251 Use `invert` to return cut off part of the input object. 2252 2253 Returns an `UnstructuredGrid` object. 2254 """ 2255 ug = self.dataset 2256 2257 ippd = vtki.new("ImplicitPolyDataDistance") 2258 ippd.SetInput(mesh.dataset) 2259 2260 if whole_cells or on_boundary: 2261 clipper = vtki.new("ExtractGeometry") 2262 clipper.SetInputData(ug) 2263 clipper.SetImplicitFunction(ippd) 2264 clipper.SetExtractInside(not invert) 2265 clipper.SetExtractBoundaryCells(False) 2266 if on_boundary: 2267 clipper.SetExtractBoundaryCells(True) 2268 clipper.SetExtractOnlyBoundaryCells(True) 2269 else: 2270 signed_dists = vtki.vtkFloatArray() 2271 signed_dists.SetNumberOfComponents(1) 2272 signed_dists.SetName("SignedDistance") 2273 for pointId in range(ug.GetNumberOfPoints()): 2274 p = ug.GetPoint(pointId) 2275 signed_dist = ippd.EvaluateFunction(p) 2276 signed_dists.InsertNextValue(signed_dist) 2277 ug.GetPointData().AddArray(signed_dists) 2278 ug.GetPointData().SetActiveScalars("SignedDistance") # NEEDED 2279 clipper = vtki.new("ClipDataSet") 2280 clipper.SetInputData(ug) 2281 clipper.SetInsideOut(not invert) 2282 clipper.SetValue(0.0) 2283 2284 clipper.Update() 2285 2286 out = UnstructuredGrid(clipper.GetOutput()) 2287 out.pipeline = utils.OperationNode("cut_with_mesh", parents=[self], c="#9e2a2b") 2288 return out
Cut a RectilinearGrid with a Mesh.
Use invert to return cut off part of the input object.
Returns an UnstructuredGrid object.
2290 def isosurface(self, value=None) -> "vedo.Mesh": 2291 """ 2292 Return a `Mesh` isosurface extracted from the object. 2293 2294 Set `value` as single float or list of values to draw the isosurface(s). 2295 """ 2296 scrange = self.dataset.GetScalarRange() 2297 2298 cf = vtki.new("ContourFilter") 2299 cf.UseScalarTreeOn() 2300 cf.SetInputData(self.dataset) 2301 cf.ComputeNormalsOn() 2302 2303 if value is None: 2304 value = (2 * scrange[0] + scrange[1]) / 3.0 2305 # print("automatic isosurface value =", value) 2306 cf.SetValue(0, value) 2307 else: 2308 if utils.is_sequence(value): 2309 cf.SetNumberOfContours(len(value)) 2310 for i, t in enumerate(value): 2311 cf.SetValue(i, t) 2312 else: 2313 cf.SetValue(0, value) 2314 2315 cf.Update() 2316 poly = cf.GetOutput() 2317 2318 out = vedo.mesh.Mesh(poly, c=None).phong() 2319 out.mapper.SetScalarRange(scrange[0], scrange[1]) 2320 2321 out.pipeline = utils.OperationNode( 2322 "isosurface", 2323 parents=[self], 2324 comment=f"#pts {out.dataset.GetNumberOfPoints()}", 2325 c="#4cc9f0:#e9c46a", 2326 ) 2327 return out
Return a Mesh isosurface extracted from the object.
Set value as single float or list of values to draw the isosurface(s).
Inherited Members
- vedo.core.PointAlgorithms
- apply_transform
- apply_transform_from_actor
- pos
- shift
- x
- y
- z
- rotate
- rotate_x
- rotate_y
- rotate_z
- reorient
- scale
- vedo.core.CommonAlgorithms
- pointdata
- celldata
- metadata
- rename
- memory_address
- memory_size
- modified
- box
- update_dataset
- bounds
- xbounds
- ybounds
- zbounds
- diagonal_size
- average_size
- center_of_mass
- copy_data_from
- inputdata
- npoints
- nvertices
- ncells
- cell_centers
- lines
- lines_as_flat_array
- mark_boundaries
- find_cells_in_bounds
- find_cells_along_line
- find_cells_along_plane
- keep_cell_types
- map_cells_to_points
- vertices
- points
- coordinates
- cells_as_flat_array
- cells
- cell_edge_neighbors
- map_points_to_cells
- resample_data_from
- interpolate_data_from
- add_ids
- gradient
- divergence
- vorticity
- probe
- compute_cell_size
- generate_random_data
- integrate_data
- write
- tomesh
- signed_distance
- unsigned_distance
- smooth_data
- compute_streamlines
- vedo.visual.MeshVisual
- follow_camera
- wireframe
- flat
- phong
- backface_culling
- render_lines_as_tubes
- frontface_culling
- backcolor
- bc
- linewidth
- lw
- linecolor
- lc
- texture
- vedo.visual.PointsVisual
- clone2d
- copy_properties_from
- color
- c
- alpha
- lut_color_at
- opacity
- force_opaque
- force_translucent
- point_size
- ps
- render_points_as_spheres
- lighting
- point_blurring
- cellcolors
- pointcolors
- cmap
- add_trail
- update_trail
- add_shadow
- update_shadows
- labels
- labels2d
- legend
- flagpole
- flagpost
- caption
