vedo.mesh

core
Mesh
Bases: MeshVisual, Points, MeshMetricsMixin
Build an instance of object Mesh derived from vedo.PointCloud.
Source code in vedo/mesh/core.py
33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510 511 512 513 514 515 516 517 518 519 520 521 522 523 524 525 526 527 528 529 530 531 532 533 534 535 536 537 538 539 540 541 542 543 544 545 546 547 548 549 550 551 552 553 554 555 556 557 558 559 560 561 562 563 564 565 566 567 568 569 570 571 572 573 574 575 576 577 578 579 580 581 582 583 584 585 586 587 588 589 590 591 592 593 594 595 596 597 598 599 600 601 602 603 604 605 606 607 608 609 610 611 612 613 614 615 616 617 618 619 620 621 622 623 624 625 626 627 628 629 630 631 632 633 634 635 636 637 638 639 640 641 642 643 644 645 646 647 648 649 650 651 652 653 654 655 656 657 658 659 660 661 662 663 664 665 666 667 668 669 670 671 672 673 674 675 676 677 678 679 680 681 682 683 684 685 686 687 688 689 690 691 692 693 694 695 696 697 698 699 700 701 702 703 704 705 706 707 708 709 710 711 712 713 714 715 716 717 718 719 720 721 722 723 724 725 726 727 728 729 730 731 732 733 734 735 736 737 738 739 740 741 742 743 744 745 746 747 748 749 750 751 752 753 754 755 756 757 758 759 760 761 762 763 764 765 766 767 768 769 770 771 772 773 774 775 776 777 778 779 780 781 782 783 784 785 786 787 788 789 790 791 792 793 794 795 796 797 798 799 800 801 802 803 804 805 806 807 808 809 810 811 812 813 814 815 816 817 818 819 820 821 822 823 824 825 826 827 828 829 830 831 832 833 834 835 836 837 838 839 840 841 842 843 844 845 846 847 848 849 850 851 852 853 854 855 856 857 858 859 860 861 862 863 864 865 866 867 868 869 870 871 872 873 874 875 876 877 878 879 880 881 882 883 884 885 886 887 888 889 890 891 892 893 894 895 896 897 898 899 900 901 902 903 904 905 906 907 908 909 910 911 912 913 914 915 916 917 918 919 920 921 922 923 924 925 926 927 928 929 930 931 932 933 934 935 936 937 938 939 940 941 942 943 944 945 946 947 948 949 950 951 952 953 954 955 956 957 958 959 960 961 962 963 964 965 966 967 968 969 970 971 972 973 974 975 976 977 978 979 980 981 982 983 984 985 986 987 988 989 990 991 992 993 994 995 996 997 998 999 1000 1001 1002 1003 1004 1005 1006 1007 1008 1009 1010 1011 1012 1013 1014 1015 1016 1017 1018 1019 1020 1021 1022 1023 1024 1025 1026 1027 1028 1029 1030 1031 1032 1033 1034 1035 1036 1037 1038 1039 1040 1041 1042 1043 1044 1045 1046 1047 1048 1049 1050 1051 1052 1053 1054 1055 1056 1057 1058 1059 1060 1061 1062 1063 1064 1065 1066 1067 1068 1069 1070 1071 1072 1073 1074 1075 1076 1077 1078 1079 1080 1081 1082 1083 1084 1085 1086 1087 1088 1089 1090 1091 1092 1093 1094 1095 1096 1097 1098 1099 1100 1101 1102 1103 1104 1105 1106 1107 1108 1109 1110 1111 1112 1113 1114 1115 1116 1117 1118 1119 1120 1121 1122 1123 1124 1125 1126 1127 1128 1129 1130 1131 1132 1133 1134 1135 1136 1137 1138 1139 1140 1141 1142 1143 1144 1145 1146 1147 1148 1149 1150 1151 1152 1153 1154 1155 1156 1157 1158 1159 1160 1161 1162 1163 1164 1165 1166 1167 1168 1169 1170 1171 1172 1173 1174 1175 1176 1177 1178 1179 1180 1181 1182 1183 1184 1185 1186 1187 1188 1189 1190 1191 1192 1193 1194 1195 1196 1197 1198 1199 1200 1201 1202 1203 1204 1205 1206 1207 1208 1209 1210 1211 1212 1213 1214 1215 1216 1217 1218 1219 1220 1221 1222 1223 1224 1225 1226 1227 1228 1229 1230 1231 1232 1233 1234 1235 1236 1237 1238 1239 1240 1241 1242 1243 1244 1245 1246 1247 1248 1249 1250 1251 1252 1253 1254 1255 1256 1257 1258 1259 1260 1261 1262 1263 1264 1265 1266 1267 1268 1269 1270 1271 1272 1273 1274 1275 1276 1277 1278 1279 1280 1281 1282 1283 1284 1285 1286 1287 1288 1289 1290 1291 1292 1293 1294 1295 1296 1297 1298 1299 1300 1301 1302 1303 1304 1305 1306 1307 1308 1309 1310 1311 1312 1313 1314 1315 1316 1317 1318 1319 1320 1321 1322 1323 1324 1325 1326 1327 1328 1329 1330 1331 1332 1333 1334 1335 1336 1337 1338 1339 1340 1341 1342 1343 1344 1345 1346 1347 1348 1349 1350 1351 1352 1353 1354 1355 1356 1357 1358 1359 1360 1361 1362 1363 1364 1365 1366 1367 1368 1369 1370 1371 1372 1373 1374 1375 1376 1377 1378 1379 1380 1381 1382 1383 1384 1385 1386 1387 1388 1389 1390 1391 1392 1393 1394 1395 1396 1397 1398 1399 1400 1401 1402 1403 1404 1405 1406 1407 1408 1409 1410 1411 1412 1413 1414 1415 1416 1417 1418 1419 1420 1421 1422 1423 1424 1425 1426 1427 1428 1429 1430 1431 1432 1433 1434 1435 1436 1437 1438 1439 1440 1441 1442 1443 1444 1445 1446 1447 1448 1449 1450 1451 1452 1453 1454 1455 1456 1457 1458 1459 1460 1461 1462 1463 1464 1465 1466 1467 1468 1469 1470 1471 1472 1473 1474 1475 1476 1477 1478 1479 1480 1481 1482 1483 1484 1485 1486 1487 1488 1489 1490 1491 1492 1493 1494 1495 1496 1497 1498 1499 1500 1501 1502 1503 1504 1505 1506 1507 1508 1509 1510 1511 1512 1513 1514 1515 1516 1517 1518 1519 1520 1521 1522 1523 1524 1525 1526 1527 1528 1529 1530 1531 1532 1533 1534 1535 1536 1537 1538 1539 1540 1541 1542 1543 1544 1545 1546 1547 1548 1549 1550 1551 1552 1553 1554 1555 1556 1557 1558 1559 1560 1561 1562 1563 1564 1565 1566 1567 1568 1569 1570 1571 1572 1573 1574 1575 1576 1577 1578 1579 1580 1581 1582 1583 1584 1585 1586 1587 1588 1589 1590 1591 1592 1593 1594 1595 1596 1597 1598 1599 1600 1601 1602 1603 1604 1605 1606 1607 1608 1609 1610 1611 1612 1613 1614 1615 1616 1617 1618 1619 1620 1621 1622 1623 1624 1625 1626 1627 1628 1629 1630 1631 1632 1633 1634 1635 1636 1637 1638 1639 1640 1641 1642 1643 1644 1645 1646 1647 1648 1649 1650 1651 1652 1653 1654 1655 1656 1657 1658 1659 1660 1661 1662 1663 1664 1665 1666 1667 1668 1669 1670 1671 1672 1673 1674 1675 1676 1677 1678 1679 1680 1681 1682 1683 1684 1685 1686 1687 1688 1689 1690 1691 1692 1693 1694 1695 1696 1697 1698 1699 1700 1701 1702 1703 1704 1705 1706 1707 1708 1709 1710 1711 1712 1713 1714 1715 1716 1717 1718 1719 1720 1721 1722 1723 1724 1725 1726 1727 1728 1729 1730 1731 1732 1733 1734 1735 1736 1737 1738 1739 1740 1741 1742 1743 1744 1745 1746 1747 1748 1749 1750 1751 1752 1753 1754 1755 1756 1757 1758 1759 1760 1761 1762 1763 1764 1765 1766 1767 1768 1769 1770 1771 1772 1773 1774 1775 1776 1777 1778 1779 1780 1781 1782 1783 1784 1785 1786 1787 1788 1789 1790 1791 1792 1793 1794 1795 1796 1797 1798 1799 1800 1801 1802 1803 1804 1805 1806 1807 1808 1809 1810 1811 1812 1813 1814 1815 1816 1817 1818 1819 1820 1821 1822 1823 1824 1825 1826 1827 1828 1829 1830 1831 1832 1833 1834 1835 1836 1837 1838 1839 1840 1841 1842 1843 1844 1845 1846 1847 1848 1849 1850 1851 1852 1853 1854 1855 1856 1857 1858 1859 1860 1861 1862 1863 1864 1865 1866 1867 1868 1869 1870 1871 1872 1873 1874 1875 1876 1877 1878 1879 1880 1881 1882 1883 1884 1885 1886 1887 1888 1889 1890 1891 1892 1893 1894 1895 1896 1897 1898 1899 1900 1901 1902 1903 1904 1905 1906 1907 1908 1909 1910 1911 1912 1913 1914 1915 1916 1917 1918 1919 1920 1921 1922 1923 1924 1925 1926 1927 1928 1929 1930 1931 1932 1933 1934 1935 1936 1937 1938 1939 1940 1941 1942 1943 1944 1945 1946 1947 1948 1949 1950 1951 1952 1953 1954 1955 1956 1957 1958 1959 1960 1961 1962 1963 1964 1965 1966 1967 1968 1969 1970 1971 1972 1973 1974 1975 1976 1977 1978 1979 1980 1981 1982 1983 1984 1985 1986 1987 1988 1989 1990 1991 1992 1993 1994 1995 1996 1997 1998 1999 2000 2001 2002 2003 2004 2005 2006 2007 2008 2009 2010 2011 2012 2013 2014 2015 2016 2017 2018 2019 2020 2021 2022 2023 2024 2025 2026 2027 2028 2029 2030 2031 2032 2033 2034 2035 2036 2037 2038 2039 2040 2041 2042 2043 2044 2045 2046 2047 2048 2049 2050 2051 2052 2053 2054 2055 2056 2057 2058 2059 2060 2061 2062 2063 2064 2065 2066 2067 2068 2069 2070 2071 2072 2073 2074 2075 2076 2077 2078 2079 2080 2081 2082 2083 2084 2085 2086 2087 2088 2089 2090 2091 2092 2093 2094 2095 2096 2097 2098 2099 2100 2101 2102 2103 2104 2105 2106 2107 2108 2109 2110 2111 2112 2113 2114 2115 2116 2117 2118 2119 2120 2121 2122 2123 2124 2125 2126 2127 2128 2129 2130 2131 2132 2133 2134 2135 2136 2137 2138 2139 2140 2141 2142 2143 2144 2145 2146 2147 2148 2149 2150 2151 2152 2153 2154 2155 2156 2157 2158 2159 2160 2161 2162 2163 2164 2165 2166 2167 2168 2169 2170 2171 2172 2173 2174 2175 2176 2177 2178 2179 2180 2181 2182 2183 2184 2185 2186 2187 2188 2189 2190 2191 2192 2193 2194 2195 2196 2197 2198 2199 2200 2201 2202 2203 2204 2205 2206 2207 2208 2209 2210 2211 2212 2213 2214 2215 2216 2217 2218 2219 2220 2221 2222 2223 2224 2225 2226 2227 2228 2229 2230 2231 2232 2233 2234 2235 2236 2237 2238 2239 2240 2241 2242 2243 2244 2245 2246 2247 2248 2249 2250 2251 2252 2253 2254 2255 2256 2257 2258 2259 2260 2261 2262 2263 2264 2265 2266 2267 2268 2269 2270 2271 2272 2273 2274 2275 2276 2277 2278 2279 2280 2281 2282 2283 2284 2285 2286 2287 2288 2289 2290 2291 2292 2293 2294 2295 2296 2297 2298 2299 2300 2301 2302 2303 2304 2305 2306 2307 2308 2309 2310 2311 2312 2313 2314 2315 2316 2317 2318 2319 2320 2321 2322 2323 2324 2325 2326 2327 2328 2329 2330 2331 2332 2333 2334 2335 2336 2337 2338 2339 2340 2341 2342 2343 2344 2345 2346 2347 2348 2349 2350 2351 2352 2353 2354 2355 2356 2357 2358 2359 2360 2361 2362 2363 2364 2365 2366 2367 2368 2369 2370 2371 2372 2373 2374 2375 2376 2377 2378 2379 2380 2381 2382 2383 2384 2385 2386 2387 2388 2389 2390 2391 2392 2393 2394 2395 2396 2397 2398 2399 2400 2401 2402 2403 2404 2405 2406 2407 2408 2409 2410 2411 2412 2413 2414 2415 2416 2417 2418 2419 2420 2421 2422 2423 2424 2425 2426 2427 2428 2429 2430 2431 2432 2433 2434 2435 2436 2437 2438 2439 2440 2441 2442 2443 2444 2445 2446 2447 2448 2449 2450 2451 2452 2453 2454 2455 2456 2457 2458 2459 2460 2461 2462 2463 2464 2465 2466 2467 2468 2469 2470 2471 2472 2473 2474 2475 2476 2477 2478 2479 2480 2481 2482 2483 2484 2485 2486 2487 2488 2489 2490 2491 2492 2493 2494 2495 2496 2497 2498 2499 2500 2501 2502 2503 2504 2505 2506 2507 2508 2509 2510 2511 2512 2513 2514 2515 2516 2517 2518 2519 2520 2521 2522 2523 2524 2525 2526 2527 2528 2529 2530 2531 2532 2533 2534 2535 2536 2537 2538 2539 2540 2541 2542 2543 2544 2545 2546 2547 2548 2549 2550 2551 2552 2553 2554 2555 2556 2557 2558 2559 2560 2561 2562 2563 2564 2565 2566 2567 2568 2569 2570 2571 2572 2573 2574 2575 2576 2577 2578 2579 2580 2581 2582 2583 2584 2585 2586 2587 2588 2589 2590 2591 2592 2593 2594 2595 2596 2597 2598 2599 2600 2601 2602 2603 2604 2605 2606 2607 2608 2609 2610 2611 2612 2613 2614 2615 2616 2617 2618 2619 2620 2621 2622 2623 2624 2625 2626 2627 2628 2629 2630 2631 2632 2633 2634 2635 2636 2637 2638 2639 2640 2641 2642 2643 2644 2645 2646 2647 2648 2649 2650 2651 2652 2653 2654 2655 2656 2657 2658 2659 2660 2661 2662 2663 2664 2665 2666 2667 2668 2669 2670 2671 2672 2673 2674 2675 2676 2677 2678 2679 2680 2681 2682 2683 2684 2685 2686 2687 2688 2689 2690 2691 2692 2693 2694 2695 2696 2697 2698 2699 2700 2701 2702 2703 2704 2705 2706 2707 2708 2709 2710 2711 2712 2713 2714 2715 2716 2717 2718 2719 2720 2721 2722 2723 2724 2725 2726 2727 2728 2729 2730 2731 2732 2733 2734 2735 2736 2737 2738 2739 2740 2741 2742 2743 2744 2745 2746 2747 2748 2749 2750 2751 2752 2753 2754 2755 2756 2757 2758 2759 2760 2761 2762 2763 2764 2765 2766 2767 2768 2769 2770 2771 2772 2773 2774 2775 2776 2777 2778 2779 2780 2781 2782 2783 2784 2785 2786 2787 2788 2789 2790 2791 2792 2793 2794 2795 2796 2797 2798 2799 2800 2801 2802 2803 2804 2805 2806 2807 2808 2809 2810 2811 2812 2813 2814 2815 2816 2817 2818 2819 | |
edges
property
Return an array containing the edges connectivity.
binarize(values=(255, 0), spacing=None, dims=None, origin=None)
Convert a Mesh into a Volume where
the interior voxels value is set to values[0] (255 by default), while
the exterior voxels value is set to values[1] (0 by default).
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
values
|
list
|
background and foreground values. |
(255, 0)
|
spacing
|
list
|
voxel spacing in x, y and z. |
None
|
dims
|
list
|
dimensions (nr. of voxels) of the output volume. |
None
|
origin
|
list
|
position in space of the (0,0,0) voxel. |
None
|
Examples:
Source code in vedo/mesh/core.py
2535 2536 2537 2538 2539 2540 2541 2542 2543 2544 2545 2546 2547 2548 2549 2550 2551 2552 2553 2554 2555 2556 2557 2558 2559 2560 2561 2562 2563 2564 2565 2566 2567 2568 2569 2570 2571 2572 2573 2574 2575 2576 2577 2578 2579 2580 2581 2582 2583 2584 2585 2586 2587 2588 2589 2590 2591 2592 2593 2594 2595 2596 2597 2598 2599 2600 2601 2602 2603 2604 2605 2606 2607 2608 2609 2610 2611 2612 2613 2614 2615 2616 2617 2618 2619 2620 2621 2622 2623 2624 2625 2626 2627 | |
boolean(operation, mesh2, method=0, tol=None)
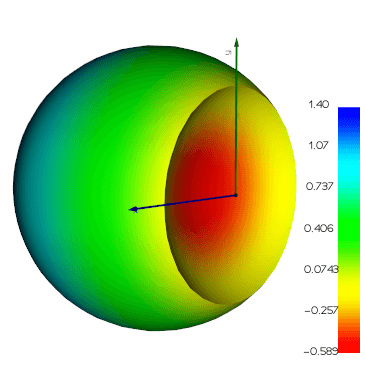
Volumetric union, intersection and subtraction of surfaces.
Use operation for the allowed operations ['plus', 'intersect', 'minus'].
Two possible algorithms are available.
Setting method to 0 (the default) uses the boolean operation algorithm
written by Cory Quammen, Chris Weigle, and Russ Taylor (https://doi.org/10.54294/216g01);
setting method to 1 will use the "loop" boolean algorithm
written by Adam Updegrove (https://doi.org/10.1016/j.advengsoft.2016.01.015).
Use tol to specify the absolute tolerance used to determine
when the distance between two points is considered to be zero (defaults to 1e-6).
Examples:
Source code in vedo/mesh/core.py
2177 2178 2179 2180 2181 2182 2183 2184 2185 2186 2187 2188 2189 2190 2191 2192 2193 2194 2195 2196 2197 2198 2199 2200 2201 2202 2203 2204 2205 2206 2207 2208 2209 2210 2211 2212 2213 2214 2215 2216 2217 2218 2219 2220 2221 2222 2223 2224 2225 2226 2227 2228 2229 2230 | |
boundaries(boundary_edges=True, manifold_edges=False, non_manifold_edges=False, feature_angle=None, return_point_ids=False, return_cell_ids=False, cell_edge=False)
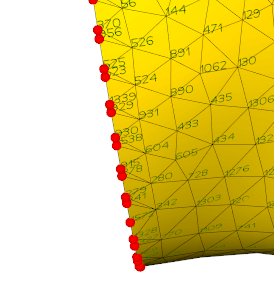
Return the boundary lines of an input mesh.
Check also vedo.core.CommonAlgorithms.mark_boundaries() method.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
boundary_edges
|
bool
|
Turn on/off the extraction of boundary edges. |
True
|
manifold_edges
|
bool
|
Turn on/off the extraction of manifold edges. |
False
|
non_manifold_edges
|
bool
|
Turn on/off the extraction of non-manifold edges. |
False
|
feature_angle
|
bool
|
Specify the min angle btw 2 faces for extracting edges. |
None
|
return_point_ids
|
bool
|
return a numpy array of point indices |
False
|
return_cell_ids
|
bool
|
return a numpy array of cell indices |
False
|
cell_edge
|
bool
|
set to |
False
|
Examples:
Source code in vedo/mesh/core.py
1415 1416 1417 1418 1419 1420 1421 1422 1423 1424 1425 1426 1427 1428 1429 1430 1431 1432 1433 1434 1435 1436 1437 1438 1439 1440 1441 1442 1443 1444 1445 1446 1447 1448 1449 1450 1451 1452 1453 1454 1455 1456 1457 1458 1459 1460 1461 1462 1463 1464 1465 1466 1467 1468 1469 1470 1471 1472 1473 1474 1475 1476 1477 1478 1479 1480 1481 1482 1483 1484 1485 1486 1487 1488 1489 1490 1491 1492 1493 1494 1495 1496 1497 1498 1499 1500 1501 1502 1503 1504 1505 1506 1507 1508 1509 1510 1511 1512 1513 1514 | |
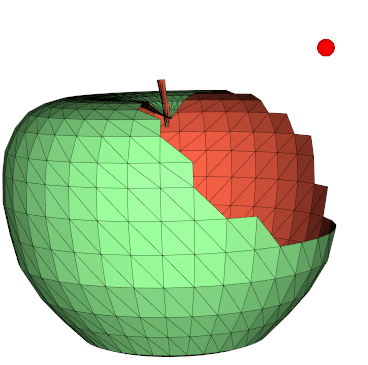
cap(return_cap=False)
Generate a "cap" on a clipped mesh, or caps sharp edges.
Examples:
See also: join(), join_segments(), slice().
Source code in vedo/mesh/core.py
377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 | |
collapse_edges(distance, iterations=1)
Collapse mesh edges so that are all above distance.
Examples:
from vedo import *
np.random.seed(2)
grid1 = Grid().add_gaussian_noise(0.8).triangulate().lw(1)
grid1.celldata['scalar'] = grid1.cell_centers().coordinates[:,1]
grid2 = grid1.clone().collapse_edges(0.1)
show(grid1, grid2, N=2, axes=1)
Source code in vedo/mesh/core.py
1158 1159 1160 1161 1162 1163 1164 1165 1166 1167 1168 1169 1170 1171 1172 1173 1174 1175 1176 1177 1178 1179 1180 1181 1182 1183 1184 1185 1186 1187 1188 1189 1190 1191 1192 1193 1194 1195 1196 1197 1198 1199 1200 1201 1202 1203 1204 | |
collide_with(mesh2, tol=0, return_bool=False)
Collide this Mesh with the input surface.
Information is stored in ContactCells1 and ContactCells2.
Source code in vedo/mesh/core.py
2425 2426 2427 2428 2429 2430 2431 2432 2433 2434 2435 2436 2437 2438 2439 2440 2441 2442 2443 2444 2445 2446 2447 2448 2449 2450 2451 2452 2453 2454 2455 2456 2457 2458 2459 2460 2461 2462 2463 2464 2465 2466 | |
compute_adjacency()
Computes the adjacency list for mesh edge-graph.
Returns:
| Type | Description |
|---|---|
list[set]
|
a list with i-th entry being the set |
list[set]
|
of indices of vertices connected by an edge to i-th vertex |
Source code in vedo/mesh/core.py
1206 1207 1208 1209 1210 1211 1212 1213 1214 1215 1216 1217 1218 1219 1220 1221 1222 1223 | |
connected_cells(index, return_ids=False)
Find all cellls connected to an input vertex specified by its index.
Source code in vedo/mesh/core.py
1619 1620 1621 1622 1623 1624 1625 1626 1627 1628 1629 1630 1631 1632 1633 1634 1635 1636 1637 1638 1639 1640 1641 1642 1643 1644 1645 1646 1647 1648 1649 1650 | |
connected_vertices(index)
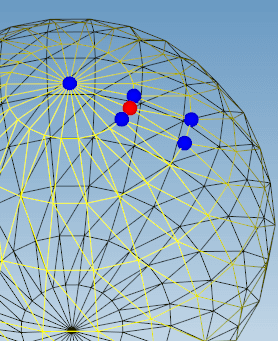
Find all vertices connected to an input vertex specified by its index.
Examples:
Source code in vedo/mesh/core.py
1563 1564 1565 1566 1567 1568 1569 1570 1571 1572 1573 1574 1575 1576 1577 1578 1579 1580 1581 1582 1583 1584 1585 1586 1587 1588 | |
contains(point, tol=1e-05)
Return True if point is inside a polydata closed surface.
Note
if you have many points to check use inside_points() instead.
Examples:
from vedo import *
s = Sphere().c('green5').alpha(0.5)
pt = [0.1, 0.2, 0.3]
print("Sphere contains", pt, s.contains(pt))
show(s, Point(pt), axes=1).close()
Source code in vedo/mesh/core.py
1331 1332 1333 1334 1335 1336 1337 1338 1339 1340 1341 1342 1343 1344 1345 1346 1347 1348 1349 1350 1351 1352 1353 1354 1355 1356 1357 | |
cut_closed_surface(origins, normals, invert=False, return_assembly=False)
Cut/clip a closed surface mesh with a collection of planes. This will produce a new closed surface by creating new polygonal faces where the input surface hits the planes.
The orientation of the polygons that form the surface is important. Polygons have a front face and a back face, and it's the back face that defines the interior or "solid" region of the closed surface. When a plane cuts through a "solid" region, a new cut face is generated, but not when a clipping plane cuts through a hole or "empty" region. This distinction is crucial when dealing with complex surfaces. Note that if a simple surface has its back faces pointing outwards, then that surface defines a hole in a potentially infinite solid.
Non-manifold surfaces should not be used with this method.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
origins
|
list
|
list of plane origins |
required |
normals
|
list
|
list of plane normals |
required |
invert
|
bool
|
invert the clipping. |
False
|
return_assembly
|
bool
|
return the cap and the clipped surfaces as a |
False
|
Examples:
from vedo import *
s = Sphere(res=50).linewidth(1)
origins = [[-0.7, 0, 0], [0, -0.6, 0]]
normals = [[-1, 0, 0], [0, -1, 0]]
s.cut_closed_surface(origins, normals)
show(s, axes=1).close()
Source code in vedo/mesh/core.py
2340 2341 2342 2343 2344 2345 2346 2347 2348 2349 2350 2351 2352 2353 2354 2355 2356 2357 2358 2359 2360 2361 2362 2363 2364 2365 2366 2367 2368 2369 2370 2371 2372 2373 2374 2375 2376 2377 2378 2379 2380 2381 2382 2383 2384 2385 2386 2387 2388 2389 2390 2391 2392 2393 2394 2395 2396 2397 2398 2399 2400 2401 2402 2403 2404 2405 2406 2407 2408 2409 2410 2411 2412 2413 2414 2415 2416 2417 2418 2419 2420 2421 2422 2423 | |
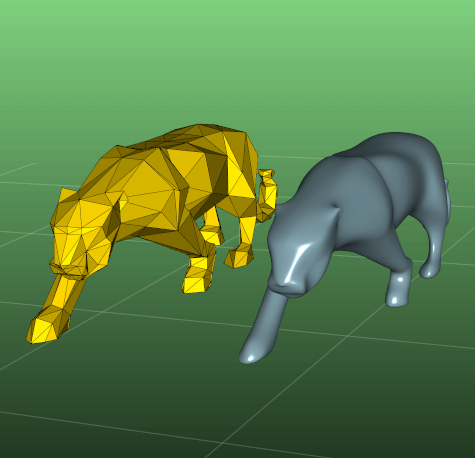
decimate(fraction=0.5, n=None, preserve_volume=True, regularization=0.0)
Downsample the number of vertices in a mesh to fraction.
This filter preserves the pointdata of the input dataset. In previous versions
of vedo, this decimation algorithm was referred to as quadric decimation.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
fraction
|
float
|
the desired target of reduction. |
0.5
|
n
|
int
|
the desired number of final points
( |
None
|
preserve_volume
|
bool
|
Decide whether to activate volume preservation which greatly reduces errors in triangle normal direction. |
True
|
regularization
|
float
|
regularize the point finding algorithm so as to have better quality mesh elements at the cost of a slightly lower precision on the geometry potentially (mostly at sharp edges). Can be useful for decimating meshes that have been triangulated on noisy data. |
0.0
|
Note
Setting fraction=0.1 leaves 10% of the original number of vertices.
Internally the VTK class
vtkQuadricDecimation
is used for this operation.
See also: decimate_binned() and decimate_pro().
Source code in vedo/mesh/core.py
842 843 844 845 846 847 848 849 850 851 852 853 854 855 856 857 858 859 860 861 862 863 864 865 866 867 868 869 870 871 872 873 874 875 876 877 878 879 880 881 882 883 884 885 886 887 888 889 890 891 892 893 894 895 896 897 898 899 900 901 902 903 904 905 | |
decimate_binned(divisions=(), use_clustering=False)
Downsample the number of vertices in a mesh.
This filter preserves the celldata of the input dataset,
if use_clustering=True also the pointdata will be preserved in the result.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
divisions
|
list
|
number of divisions along x, y and z axes. |
()
|
auto_adjust
|
bool
|
if True, the number of divisions is automatically adjusted to create more uniform cells. |
required |
use_clustering
|
bool
|
use vtkQuadricClustering instead of vtkBinnedDecimation. |
False
|
See also: decimate() and decimate_pro().
Source code in vedo/mesh/core.py
993 994 995 996 997 998 999 1000 1001 1002 1003 1004 1005 1006 1007 1008 1009 1010 1011 1012 1013 1014 1015 1016 1017 1018 1019 1020 1021 1022 1023 1024 1025 1026 1027 1028 1029 1030 1031 1032 1033 1034 1035 1036 1037 | |
decimate_pro(fraction=0.5, n=None, preserve_topology=True, preserve_boundaries=True, splitting=False, splitting_angle=75, feature_angle=0, inflection_point_ratio=10, vertex_degree=0)
Downsample the number of vertices in a mesh to fraction.
This filter preserves the pointdata of the input dataset.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
fraction
|
float
|
The desired target of reduction.
Setting |
0.5
|
n
|
int
|
the desired number of final points ( |
None
|
preserve_topology
|
bool
|
If on, mesh splitting and hole elimination will not occur. This may limit the maximum reduction that may be achieved. |
True
|
preserve_boundaries
|
bool
|
Turn on/off the deletion of vertices on the boundary of a mesh. Control whether mesh boundaries are preserved during decimation. |
True
|
feature_angle
|
float
|
Specify the angle that defines a feature. This angle is used to define what an edge is (i.e., if the surface normal between two adjacent triangles is >= FeatureAngle, an edge exists). |
0
|
splitting
|
bool
|
Turn on/off the splitting of the mesh at corners, along edges, at non-manifold points, or anywhere else a split is required. Turning splitting off will better preserve the original topology of the mesh, but you may not obtain the requested reduction. |
False
|
splitting_angle
|
float
|
Specify the angle that defines a sharp edge.
This angle is used to control the splitting of the mesh.
A split line exists when the surface normals between two edge connected triangles
are >= |
75
|
inflection_point_ratio
|
float
|
An inflection point occurs when the ratio of reduction error between two iterations
is greater than or equal to the |
10
|
vertex_degree
|
int
|
If the number of triangles connected to a vertex exceeds it then the vertex will be split. |
0
|
Note
Setting fraction=0.1 leaves 10% of the original number of vertices
See also
decimate() and decimate_binned().
Source code in vedo/mesh/core.py
907 908 909 910 911 912 913 914 915 916 917 918 919 920 921 922 923 924 925 926 927 928 929 930 931 932 933 934 935 936 937 938 939 940 941 942 943 944 945 946 947 948 949 950 951 952 953 954 955 956 957 958 959 960 961 962 963 964 965 966 967 968 969 970 971 972 973 974 975 976 977 978 979 980 981 982 983 984 985 986 987 988 989 990 991 | |
delete_cells(ids)
Remove cells from the mesh object by their ID.
Points (vertices) are not removed (you may use clean() to remove those).
Source code in vedo/mesh/core.py
1115 1116 1117 1118 1119 1120 1121 1122 1123 1124 1125 1126 1127 1128 1129 1130 1131 | |
delete_cells_by_point_index(indices)
Delete a list of vertices identified by any of their vertex index.
See also delete_cells().
Examples:
Source code in vedo/mesh/core.py
1133 1134 1135 1136 1137 1138 1139 1140 1141 1142 1143 1144 1145 1146 1147 1148 1149 1150 1151 1152 1153 1154 1155 1156 | |
extract_cells(ids)
Extract a subset of cells from a mesh and return it as a new mesh.
Source code in vedo/mesh/core.py
1590 1591 1592 1593 1594 1595 1596 1597 1598 1599 1600 1601 1602 1603 1604 1605 1606 1607 1608 1609 1610 1611 1612 1613 1614 1615 1616 1617 | |
extract_largest_region()
Extract the largest connected part of a mesh and discard all the smaller pieces.
Examples:
Source code in vedo/mesh/core.py
2154 2155 2156 2157 2158 2159 2160 2161 2162 2163 2164 2165 2166 2167 2168 2169 2170 2171 2172 2173 2174 2175 | |
extrude(zshift=1.0, direction=(), rotation=0.0, dr=0.0, cap=True, res=1)
Sweep a polygonal data creating a "skirt" from free edges and lines, and lines from vertices. The input dataset is swept around the z-axis to create new polygonal primitives. For example, sweeping a line results in a cylindrical shell, and sweeping a circle creates a torus.
You can control whether the sweep of a 2D object (i.e., polygon or triangle strip) is capped with the generating geometry. Also, you can control the angle of rotation, and whether translation along the z-axis is performed along with the rotation. (Translation is useful for creating "springs"). You also can adjust the radius of the generating geometry using the "dR" keyword.
The skirt is generated by locating certain topological features. Free edges (edges of polygons or triangle strips only used by one polygon or triangle strips) generate surfaces. This is true also of lines or polylines. Vertices generate lines.
This filter can be used to model axisymmetric objects like cylinders, bottles, and wine glasses; or translational/rotational symmetric objects like springs or corkscrews.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
zshift
|
float
|
shift along z axis. |
1.0
|
direction
|
list
|
extrusion direction in the xy plane. note that zshift is forced to be the 3rd component of direction, which is therefore ignored. |
()
|
rotation
|
float
|
set the angle of rotation. |
0.0
|
dr
|
float
|
set the radius variation in absolute units. |
0.0
|
cap
|
bool
|
enable or disable capping. |
True
|
res
|
int
|
set the resolution of the generating geometry. |
1
|
Warning
Some polygonal objects have no free edges (e.g., sphere). When swept, this will result in two separate surfaces if capping is on, or no surface if capping is off.
Examples:
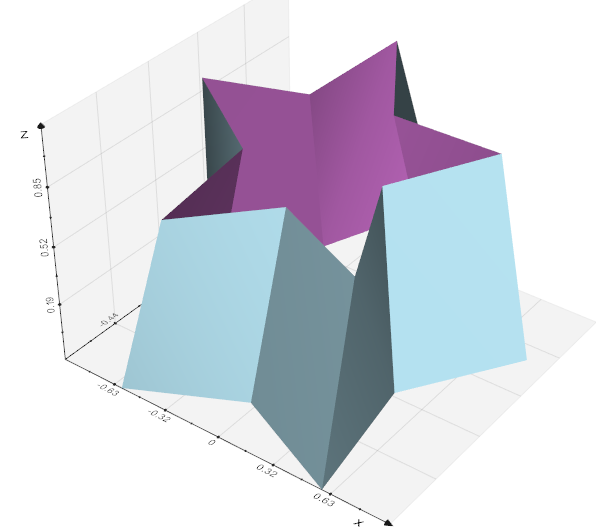
Source code in vedo/mesh/core.py
1833 1834 1835 1836 1837 1838 1839 1840 1841 1842 1843 1844 1845 1846 1847 1848 1849 1850 1851 1852 1853 1854 1855 1856 1857 1858 1859 1860 1861 1862 1863 1864 1865 1866 1867 1868 1869 1870 1871 1872 1873 1874 1875 1876 1877 1878 1879 1880 1881 1882 1883 1884 1885 1886 1887 1888 1889 1890 1891 1892 1893 1894 1895 1896 1897 1898 1899 1900 1901 1902 1903 1904 1905 1906 1907 1908 1909 | |
extrude_and_trim_with(surface, direction=(), strategy='all', cap=True, cap_strategy='max')
Extrude a Mesh and trim it with an input surface mesh.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
surface
|
Mesh
|
the surface mesh to trim with. |
required |
direction
|
list
|
extrusion direction in the xy plane. |
()
|
strategy
|
str
|
either "boundary_edges" or "all_edges". |
'all'
|
cap
|
bool
|
enable or disable capping. |
True
|
cap_strategy
|
str
|
either "intersection", "minimum_distance", "maximum_distance", "average_distance". |
'max'
|
The input Mesh is swept along a specified direction forming a "skirt" from the boundary edges 2D primitives (i.e., edges used by only one polygon); and/or from vertices and lines. The extent of the sweeping is limited by a second input: defined where the sweep intersects a user-specified surface.
Capping of the extrusion can be enabled. In this case the input, generating primitive is copied inplace as well as to the end of the extrusion skirt. (See warnings below on what happens if the intersecting sweep does not intersect, or partially intersects the trim surface.)
Note that this method operates in two fundamentally different modes based on the extrusion strategy. If the strategy is "boundary_edges", then only the boundary edges of the input's 2D primitives are extruded (verts and lines are extruded to generate lines and quads). However, if the extrusions strategy is "all_edges", then every edge of the 2D primitives is used to sweep out a quadrilateral polygon (again verts and lines are swept to produce lines and quads).
Warning
The extrusion direction is assumed to define an infinite line. The intersection with the trim surface is along a ray from the - to + direction, however only the first intersection is taken. Some polygonal objects have no free edges (e.g., sphere). When swept, this will result in two separate surfaces if capping is on and "boundary_edges" enabled, or no surface if capping is off and "boundary_edges" is enabled. If all the extrusion lines emanating from an extruding primitive do not intersect the trim surface, then no output for that primitive will be generated. In extreme cases, it is possible that no output whatsoever will be generated.
Examples:
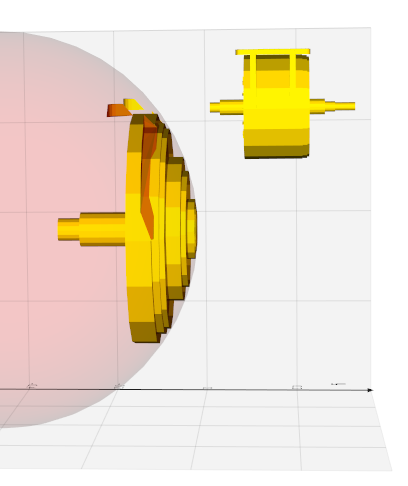
from vedo import *
sphere = Sphere([-1,0,4]).rotate_x(25).wireframe().color('red5')
circle = Circle([0,0,0], r=2, res=100).color('b6')
extruded_circle = circle.extrude_and_trim_with(
sphere,
direction=[0,-0.2,1],
strategy="bound",
cap=True,
cap_strategy="intersection",
)
circle.lw(3).color("tomato").shift(dz=-0.1)
show(circle, sphere, extruded_circle, axes=1).close()
Source code in vedo/mesh/core.py
1963 1964 1965 1966 1967 1968 1969 1970 1971 1972 1973 1974 1975 1976 1977 1978 1979 1980 1981 1982 1983 1984 1985 1986 1987 1988 1989 1990 1991 1992 1993 1994 1995 1996 1997 1998 1999 2000 2001 2002 2003 2004 2005 2006 2007 2008 2009 2010 2011 2012 2013 2014 2015 2016 2017 2018 2019 2020 2021 2022 2023 2024 2025 2026 2027 2028 2029 2030 2031 2032 2033 2034 2035 2036 2037 2038 2039 2040 2041 2042 2043 2044 2045 2046 2047 2048 2049 2050 2051 2052 2053 2054 2055 2056 2057 2058 2059 2060 2061 2062 2063 2064 2065 2066 2067 2068 2069 | |
extrude_linear(shift=1.0, direction=(0, 0, 1), cap=True, use_normal=False)
Linearly extrude polygonal data using vtkLinearExtrusionFilter.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
shift
|
float
|
Extrusion scale factor. With vector extrusion, the final displacement is
|
1.0
|
direction
|
list
|
Extrusion vector. Ignored when |
(0, 0, 1)
|
cap
|
bool
|
Enable or disable capping. |
True
|
use_normal
|
bool
|
If |
False
|
Examples:
Source code in vedo/mesh/core.py
1911 1912 1913 1914 1915 1916 1917 1918 1919 1920 1921 1922 1923 1924 1925 1926 1927 1928 1929 1930 1931 1932 1933 1934 1935 1936 1937 1938 1939 1940 1941 1942 1943 1944 1945 1946 1947 1948 1949 1950 1951 1952 1953 1954 1955 1956 1957 1958 1959 1960 1961 | |
fill_holes(size=None)
Identifies and fills holes in the input mesh. Holes are identified by locating boundary edges, linking them together into loops, and then triangulating the resulting loops.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
size
|
float
|
Approximate limit to the size of the hole that can be filled. |
None
|
Examples:
Source code in vedo/mesh/core.py
1301 1302 1303 1304 1305 1306 1307 1308 1309 1310 1311 1312 1313 1314 1315 1316 1317 1318 1319 1320 1321 1322 1323 1324 1325 1326 1327 1328 1329 | |
find_adjacent_vertices(index, depth=1, adjacency_list=None)
Computes the ball of radius n in the mesh' edge-graph metric
centered at vertex index.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
index
|
int
|
index of the vertex |
required |
depth
|
int
|
depth of the search in the edge-graph metric. |
1
|
Returns:
| Type | Description |
|---|---|
set
|
the set of indices of the vertices which are at most |
Source code in vedo/mesh/core.py
1225 1226 1227 1228 1229 1230 1231 1232 1233 1234 1235 1236 1237 1238 1239 1240 1241 1242 1243 1244 1245 1246 1247 1248 1249 1250 1251 1252 1253 1254 | |
generate_random_points(n, min_radius=0.0)
Generate n uniformly distributed random points
inside the polygonal mesh.
A new point data array is added to the output points called "OriginalCellID" which contains the index of the cell ID in which the point was generated.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
n
|
int
|
number of points to generate. |
required |
min_radius
|
float
|
impose a minimum distance between points.
If |
0.0
|
Returns a vedo.Points object.
Note
Consider using points.probe(msh) or
points.interpolate_data_from(msh)
to interpolate existing mesh data onto the new points.
Examples:
from vedo import *
msh = Mesh(dataurl + "panther.stl").lw(2)
pts = msh.generate_random_points(20000, min_radius=0.5)
print("Original cell ids:", pts.pointdata["OriginalCellID"])
show(pts, msh, axes=1).close()
Source code in vedo/mesh/core.py
1039 1040 1041 1042 1043 1044 1045 1046 1047 1048 1049 1050 1051 1052 1053 1054 1055 1056 1057 1058 1059 1060 1061 1062 1063 1064 1065 1066 1067 1068 1069 1070 1071 1072 1073 1074 1075 1076 1077 1078 1079 1080 1081 1082 1083 1084 1085 1086 1087 1088 1089 1090 1091 1092 1093 1094 1095 1096 1097 1098 1099 1100 1101 1102 1103 1104 1105 1106 1107 1108 1109 1110 1111 1112 1113 | |
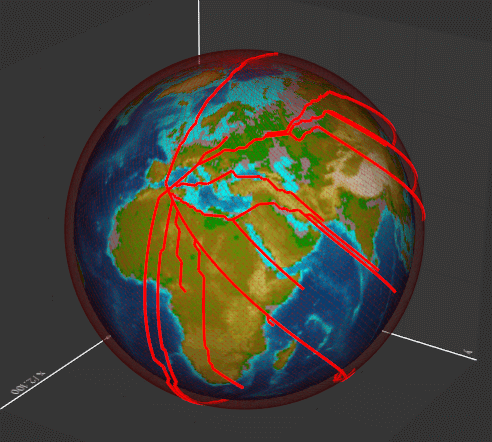
geodesic(start, end)
Dijkstra algorithm to compute the geodesic line. Takes as input a polygonal mesh and performs a single source shortest path calculation.
The output mesh contains the array "VertexIDs" that contains the ordered list of vertices traversed to get from the start vertex to the end vertex.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
start
|
(int, list)
|
start vertex index or close point |
required |
end
|
(int, list)
|
end vertex index or close point |
required |
Examples:
Source code in vedo/mesh/core.py
2468 2469 2470 2471 2472 2473 2474 2475 2476 2477 2478 2479 2480 2481 2482 2483 2484 2485 2486 2487 2488 2489 2490 2491 2492 2493 2494 2495 2496 2497 2498 2499 2500 2501 2502 2503 2504 2505 2506 2507 2508 2509 2510 2511 2512 2513 2514 2515 2516 2517 2518 2519 2520 2521 2522 2523 2524 2525 2526 2527 2528 2529 2530 | |
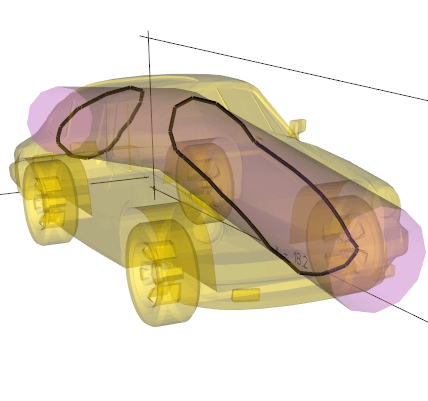
imprint(loopline, tol=0.01)
Imprint the contact surface of one object onto another surface.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
loopline
|
Line
|
a Line object to be imprinted onto the mesh. |
required |
tol
|
float
|
projection tolerance which controls how close the imprint surface must be to the target. |
0.01
|
Examples:
from vedo import *
grid = Grid()#.triangulate()
circle = Circle(r=0.3, res=24).pos(0.11,0.12)
line = Line(circle, closed=True, lw=4, c='r4')
grid.imprint(line)
show(grid, line, axes=1).close()
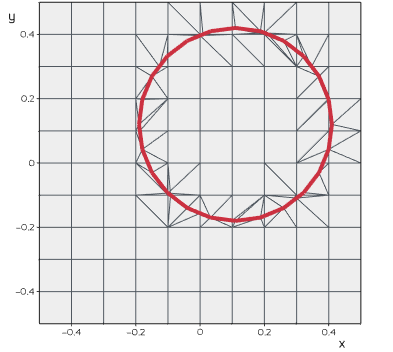
Source code in vedo/mesh/core.py
1516 1517 1518 1519 1520 1521 1522 1523 1524 1525 1526 1527 1528 1529 1530 1531 1532 1533 1534 1535 1536 1537 1538 1539 1540 1541 1542 1543 1544 1545 1546 1547 1548 1549 1550 1551 1552 1553 1554 1555 1556 1557 1558 1559 1560 1561 | |
inside_points(pts, invert=False, tol=1e-05, return_ids=False)
Return the point cloud that is inside mesh surface as a new Points object.
If return_ids is True a list of IDs is returned and in addition input points are marked by a pointdata array named "IsInside".
Examples:
print(pts.pointdata["IsInside"])
Examples:
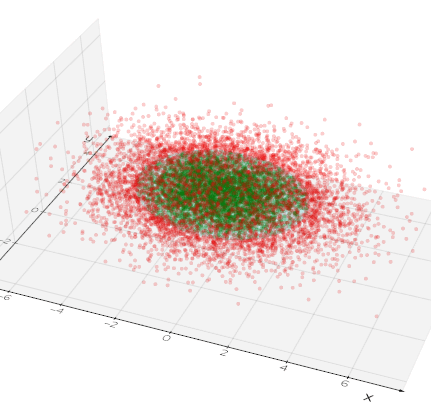
Source code in vedo/mesh/core.py
1359 1360 1361 1362 1363 1364 1365 1366 1367 1368 1369 1370 1371 1372 1373 1374 1375 1376 1377 1378 1379 1380 1381 1382 1383 1384 1385 1386 1387 1388 1389 1390 1391 1392 1393 1394 1395 1396 1397 1398 1399 1400 1401 1402 1403 1404 1405 1406 1407 1408 1409 1410 1411 1412 1413 | |
intersect_with(mesh2, tol=1e-06)
Intersect this Mesh with the input surface to return a set of lines.
Examples:
Source code in vedo/mesh/core.py
2232 2233 2234 2235 2236 2237 2238 2239 2240 2241 2242 2243 2244 2245 2246 2247 2248 2249 2250 2251 2252 2253 | |
intersect_with_line(p0, p1=None, return_ids=False, tol=0)
Return the list of points intersecting the mesh
along the segment defined by two points p0 and p1.
Use return_ids to return the cell ids along with point coords
Examples:
from vedo import *
s = Spring()
pts = s.intersect_with_line([0,0,0], [1,0.1,0])
ln = Line([0,0,0], [1,0.1,0], c='blue')
ps = Points(pts, r=10, c='r')
show(s, ln, ps, bg='white').close()
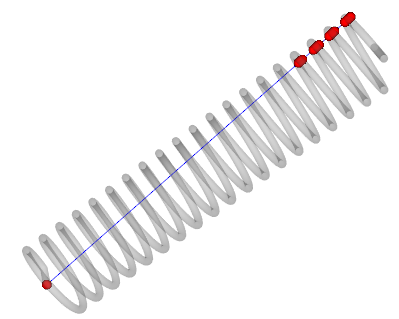
Source code in vedo/mesh/core.py
2255 2256 2257 2258 2259 2260 2261 2262 2263 2264 2265 2266 2267 2268 2269 2270 2271 2272 2273 2274 2275 2276 2277 2278 2279 2280 2281 2282 2283 2284 2285 2286 2287 2288 2289 2290 2291 2292 2293 2294 2295 2296 2297 2298 2299 2300 2301 2302 2303 | |
intersect_with_plane(origin=(0, 0, 0), normal=(1, 0, 0))
Intersect this Mesh with a plane to return a set of lines.
Examples:
from vedo import *
sph = Sphere()
mi = sph.clone().intersect_with_plane().join()
print(mi.lines)
show(sph, mi, axes=1).close()
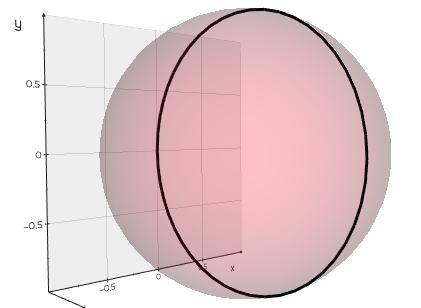
Source code in vedo/mesh/core.py
2305 2306 2307 2308 2309 2310 2311 2312 2313 2314 2315 2316 2317 2318 2319 2320 2321 2322 2323 2324 2325 2326 2327 2328 2329 2330 2331 2332 2333 2334 2335 2336 2337 2338 | |
isobands(n=10, vmin=None, vmax=None)
Return a new Mesh representing the isobands of the active scalars.
This is a new mesh where the scalar is now associated to cell faces and
used to colorize the mesh.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
n
|
int
|
number of isobands in the range |
10
|
vmin
|
float
|
minimum of the range |
None
|
vmax
|
float
|
maximum of the range |
None
|
Examples:
Source code in vedo/mesh/core.py
1722 1723 1724 1725 1726 1727 1728 1729 1730 1731 1732 1733 1734 1735 1736 1737 1738 1739 1740 1741 1742 1743 1744 1745 1746 1747 1748 1749 1750 1751 1752 1753 1754 1755 1756 1757 1758 1759 1760 1761 1762 1763 1764 1765 1766 1767 1768 1769 1770 1771 1772 1773 1774 1775 1776 1777 1778 1779 1780 1781 1782 | |
isolines(n=10, vmin=None, vmax=None)
Return a new Mesh representing the isolines of the active scalars.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
n
|
(int, list)
|
number of isolines in the range, a list of specific values can also be passed. |
10
|
vmin
|
float
|
minimum of the range |
None
|
vmax
|
float
|
maximum of the range |
None
|
Examples:
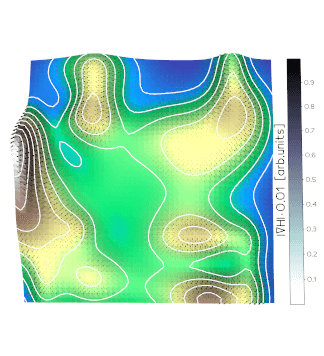
Source code in vedo/mesh/core.py
1784 1785 1786 1787 1788 1789 1790 1791 1792 1793 1794 1795 1796 1797 1798 1799 1800 1801 1802 1803 1804 1805 1806 1807 1808 1809 1810 1811 1812 1813 1814 1815 1816 1817 1818 1819 1820 1821 1822 1823 1824 1825 1826 1827 1828 1829 1830 1831 | |
join(polys=True, reset=False)
Generate triangle strips and/or polylines from input polygons, triangle strips, and lines.
Input polygons are assembled into triangle strips only if they are triangles; other types of polygons are passed through to the output and not stripped. Use mesh.triangulate() to triangulate non-triangular polygons prior to running this filter if you need to strip all the data.
Also note that if triangle strips or polylines are present in the input they are passed through and not joined nor extended. If you wish to strip these use mesh.triangulate() to fragment the input into triangles and lines prior to applying join().
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
polys
|
bool
|
polygonal segments will be joined if they are contiguous |
True
|
reset
|
bool
|
reset points ordering |
False
|
Warning
If triangle strips or polylines exist in the input data they will be passed through to the output data. This filter will only construct triangle strips if triangle polygons are available; and will only construct polylines if lines are available.
Examples:
from vedo import *
c1 = Cylinder(pos=(0,0,0), r=2, height=3, axis=(1,.0,0), alpha=.1).triangulate()
c2 = Cylinder(pos=(0,0,2), r=1, height=2, axis=(0,.3,1), alpha=.1).triangulate()
intersect = c1.intersect_with(c2).join(reset=True)
spline = Spline(intersect).c('blue').lw(5)
show(c1, c2, spline, intersect.labels('id'), axes=1).close()
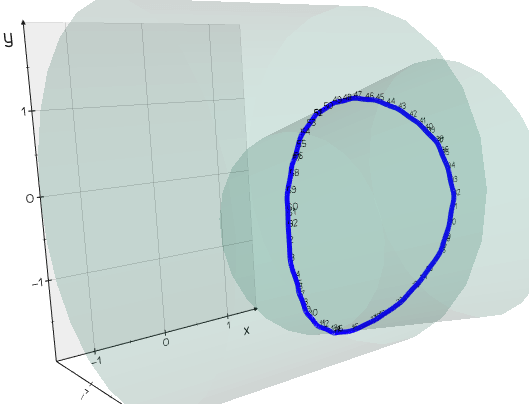
Source code in vedo/mesh/core.py
435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 | |
join_segments(closed=True, tol=0.001)
Join line segments into contiguous lines.
Useful to call with triangulate() method.
Returns:
| Type | Description |
|---|---|
list
|
list of |
Examples:
from vedo import *
msh = Torus().alpha(0.1).wireframe()
intersection = msh.intersect_with_plane(normal=[1,1,1]).c('purple5')
slices = [s.triangulate() for s in intersection.join_segments()]
show(msh, intersection, merge(slices), axes=1, viewup='z')
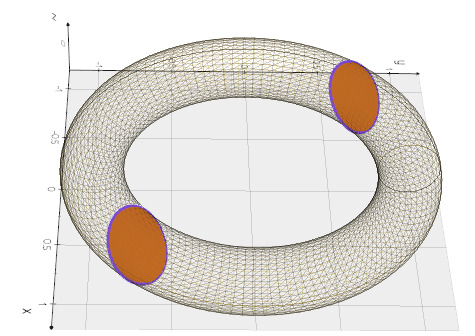
Source code in vedo/mesh/core.py
501 502 503 504 505 506 507 508 509 510 511 512 513 514 515 516 517 518 519 520 521 522 523 524 525 526 527 528 529 530 531 532 533 534 535 536 537 538 539 540 541 542 543 544 545 546 547 548 549 550 551 552 553 554 555 556 557 558 559 560 561 562 563 564 565 566 567 568 | |
join_with_strips(b1, closed=True)
Join booundary lines by creating a triangle strip between them.
Examples:
from vedo import *
m1 = Cylinder(cap=False).boundaries()
m2 = Cylinder(cap=False).boundaries().pos(0.2,0,1)
strips = m1.join_with_strips(m2)
show(m1, m2, strips, axes=1).close()
Source code in vedo/mesh/core.py
570 571 572 573 574 575 576 577 578 579 580 581 582 583 584 585 586 587 588 589 590 591 592 593 594 595 596 597 598 599 600 601 602 603 604 605 606 607 608 609 610 611 612 613 614 615 616 617 618 619 620 621 622 623 624 625 626 627 628 | |
laplacian_diffusion(array_name, dt, num_steps)
Apply a diffusion process to a scalar array defined on the points of a mesh.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
array_name
|
str
|
name of the array to diffuse. |
required |
dt
|
float
|
time step. |
required |
num_steps
|
int
|
number of iterations. |
required |
Source code in vedo/mesh/core.py
721 722 723 724 725 726 727 728 729 730 731 732 733 734 735 736 737 738 739 740 741 742 743 744 745 746 747 748 749 750 751 752 753 754 755 756 757 758 759 760 761 762 763 764 765 766 767 768 769 770 771 772 773 774 775 776 777 778 779 780 781 782 783 784 785 786 787 788 789 790 | |
remove_all_lines()
Remove all line elements from the mesh.
Source code in vedo/mesh/core.py
646 647 648 649 | |
reverse(cells=True, normals=False)
Reverse the order of polygonal cells and/or reverse the direction of point and cell normals.
Two flags are used to control these operations
cells=Truereverses the order of the indices in the cell connectivity list. If cell is a list of IDs only those cells will be reversed.normals=Truereverses the normals by multiplying the normal vector by -1 (both point and cell normals, if present).
Source code in vedo/mesh/core.py
287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 | |
shrink(fraction=0.85)
Shrink the triangle polydata in the representation of the input mesh.
Examples:
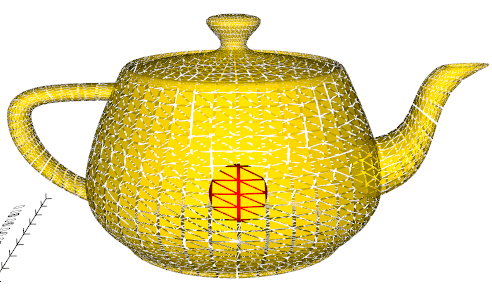
Source code in vedo/mesh/core.py
359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 | |
signed_distance(bounds=None, dims=(20, 20, 20), invert=False, max_radius=None)
Compute the Volume object whose voxels contains
the signed distance from the mesh.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
bounds
|
list
|
bounds of the output volume |
None
|
dims
|
list
|
dimensions (nr. of voxels) of the output volume |
(20, 20, 20)
|
invert
|
bool
|
flip the sign |
False
|
max_radius
|
float
|
ignored for meshes (only valid for point clouds); kept for API uniformity |
None
|
Examples:
Source code in vedo/mesh/core.py
2629 2630 2631 2632 2633 2634 2635 2636 2637 2638 2639 2640 2641 2642 2643 2644 2645 2646 2647 2648 2649 2650 2651 2652 2653 2654 2655 2656 2657 2658 2659 2660 2661 2662 2663 2664 2665 2666 2667 2668 2669 2670 2671 2672 2673 2674 2675 2676 2677 2678 2679 2680 2681 2682 2683 2684 2685 2686 2687 2688 2689 2690 | |
silhouette(direction=None, border_edges=True, feature_angle=False)
Return a new line Mesh which corresponds to the outer silhouette
of the input as seen along a specified direction, this can also be
a vtkCamera object.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
direction
|
list
|
viewpoint direction vector.
If |
None
|
border_edges
|
bool
|
enable or disable generation of border edges |
True
|
feature_angle
|
float
|
minimal angle for sharp edges detection.
If set to |
False
|
Examples:
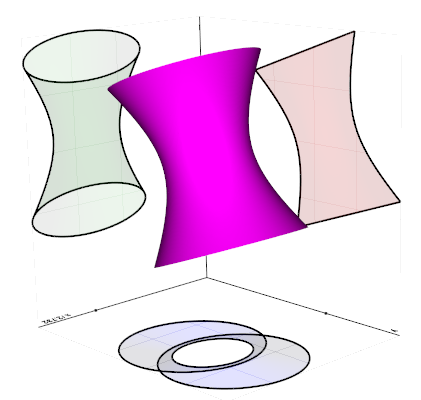
Source code in vedo/mesh/core.py
1652 1653 1654 1655 1656 1657 1658 1659 1660 1661 1662 1663 1664 1665 1666 1667 1668 1669 1670 1671 1672 1673 1674 1675 1676 1677 1678 1679 1680 1681 1682 1683 1684 1685 1686 1687 1688 1689 1690 1691 1692 1693 1694 1695 1696 1697 1698 1699 1700 1701 1702 1703 1704 1705 1706 1707 1708 1709 1710 1711 1712 1713 1714 1715 1716 1717 1718 1719 1720 | |
slice(origin=(0, 0, 0), normal=(1, 0, 0))
Slice a mesh with a plane and fill the contour.
Examples:
from vedo import *
msh = Mesh(dataurl+"bunny.obj").alpha(0.1).wireframe()
mslice = msh.slice(normal=[0,1,0.3], origin=[0,0.16,0])
mslice.c('purple5')
show(msh, mslice, axes=1)
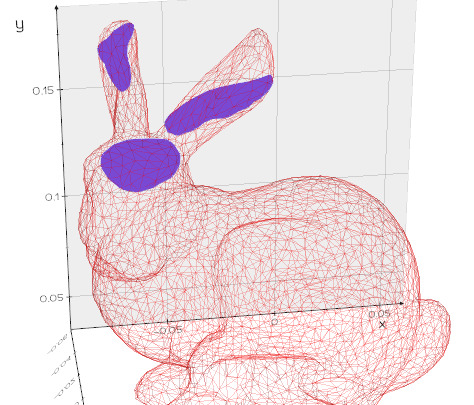
See also: join(), join_segments(), cap(), cut_with_plane().
Source code in vedo/mesh/core.py
651 652 653 654 655 656 657 658 659 660 661 662 663 664 665 666 667 668 669 670 671 672 673 674 675 | |
smooth(niter=15, pass_band=0.1, edge_angle=15, feature_angle=60, boundary=False)
Adjust mesh point positions using the so-called "Windowed Sinc" method.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
niter
|
int
|
number of iterations. |
15
|
pass_band
|
float
|
set the pass_band value for the windowed sinc filter. |
0.1
|
edge_angle
|
float
|
edge angle to control smoothing along edges (either interior or boundary). |
15
|
feature_angle
|
float
|
specifies the feature angle for sharp edge identification. |
60
|
boundary
|
bool
|
specify if boundary should also be smoothed or kept unmodified |
False
|
Examples:

Source code in vedo/mesh/core.py
1256 1257 1258 1259 1260 1261 1262 1263 1264 1265 1266 1267 1268 1269 1270 1271 1272 1273 1274 1275 1276 1277 1278 1279 1280 1281 1282 1283 1284 1285 1286 1287 1288 1289 1290 1291 1292 1293 1294 1295 1296 1297 1298 1299 | |
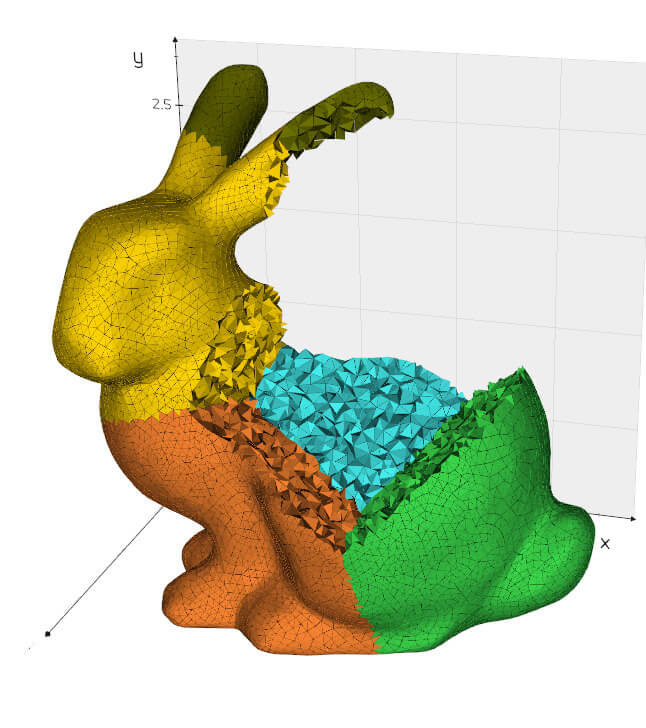
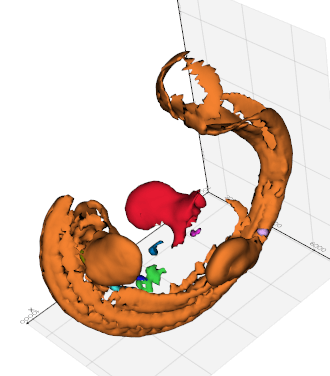
split(maxdepth=1000, flag=False, must_share_edge=False, sort_by_area=True)
Split a mesh by connectivity and order the pieces by increasing area.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
maxdepth
|
int
|
only consider this maximum number of mesh parts. |
1000
|
flag
|
bool
|
if set to True return the same single object, but add a "RegionId" array to flag the mesh subparts |
False
|
must_share_edge
|
bool
|
if True, mesh regions that only share single points will be split. |
False
|
sort_by_area
|
bool
|
if True, sort the mesh parts by decreasing area. |
True
|
Examples:

Source code in vedo/mesh/core.py
2071 2072 2073 2074 2075 2076 2077 2078 2079 2080 2081 2082 2083 2084 2085 2086 2087 2088 2089 2090 2091 2092 2093 2094 2095 2096 2097 2098 2099 2100 2101 2102 2103 2104 2105 2106 2107 2108 2109 2110 2111 2112 2113 2114 2115 2116 2117 2118 2119 2120 2121 2122 2123 2124 2125 2126 2127 2128 2129 2130 2131 2132 2133 2134 2135 2136 2137 2138 2139 2140 2141 2142 2143 2144 2145 2146 2147 2148 2149 2150 2151 2152 | |
split_polylines()
Split polylines into separate segments.
Source code in vedo/mesh/core.py
630 631 632 633 634 635 636 637 638 639 640 641 642 643 644 | |
subdivide(n=1, method=0, mel=None)
Increase the number of vertices of a surface mesh.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
n
|
int
|
number of subdivisions. |
1
|
method
|
int
|
Loop(0), Linear(1), Adaptive(2), Butterfly(3), Centroid(4) |
0
|
mel
|
float
|
Maximum Edge Length (applicable to Adaptive method only). |
None
|
Source code in vedo/mesh/core.py
792 793 794 795 796 797 798 799 800 801 802 803 804 805 806 807 808 809 810 811 812 813 814 815 816 817 818 819 820 821 822 823 824 825 826 827 828 829 830 831 832 833 834 835 836 837 838 839 840 | |
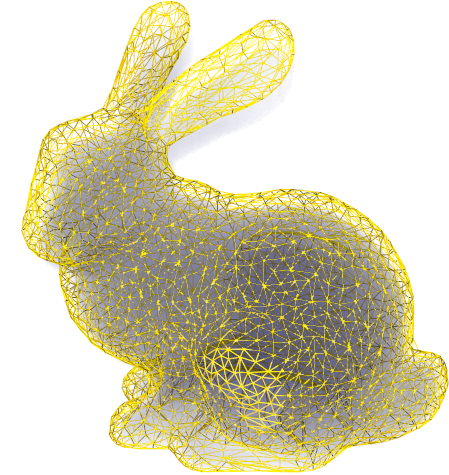
tetralize(side=0.02, nmax=300000, gap=None, subsample=False, uniform=True, seed=0, debug=False)
Tetralize a closed polygonal mesh. Return a TetMesh.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
side
|
float
|
desired side of the single tetras as fraction of the bounding box diagonal. Typical values are in the range (0.01 - 0.03) |
0.02
|
nmax
|
int
|
maximum random numbers to be sampled in the bounding box |
300000
|
gap
|
float
|
keep this minimum distance from the surface, if None an automatic choice is made. |
None
|
subsample
|
bool
|
subsample input surface, the geometry might be affected (the number of original faces reduceed), but higher tet quality might be obtained. |
False
|
uniform
|
bool
|
generate tets more uniformly packed in the interior of the mesh |
True
|
seed
|
int
|
random number generator seed |
0
|
debug
|
bool
|
show an intermediate plot with sampled points |
False
|
Examples:
Source code in vedo/mesh/core.py
2692 2693 2694 2695 2696 2697 2698 2699 2700 2701 2702 2703 2704 2705 2706 2707 2708 2709 2710 2711 2712 2713 2714 2715 2716 2717 2718 2719 2720 2721 2722 2723 2724 2725 2726 2727 2728 2729 2730 2731 2732 2733 2734 2735 2736 2737 2738 2739 2740 2741 2742 2743 2744 2745 2746 2747 2748 2749 2750 2751 2752 2753 2754 2755 2756 2757 2758 2759 2760 2761 2762 2763 2764 2765 2766 2767 2768 2769 2770 2771 2772 2773 2774 2775 2776 2777 2778 2779 2780 2781 2782 2783 2784 2785 2786 2787 2788 2789 2790 2791 2792 2793 2794 2795 2796 2797 2798 2799 2800 2801 2802 2803 2804 2805 2806 2807 2808 2809 2810 2811 2812 2813 2814 2815 2816 2817 2818 2819 | |
to_reeb_graph(field_id=0)
Convert the mesh into a Reeb graph. The Reeb graph is a topological structure that captures the evolution of the level sets of a scalar field.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
field_id
|
int
|
the id of the scalar field to use. |
0
|
Examples:
from vedo import *
mesh = Mesh("https://discourse.paraview.org/uploads/short-url/qVuZ1fiRjwhE1qYtgGE2HGXybgo.stl")
mesh.rotate_x(10).rotate_y(15).alpha(0.5)
mesh.pointdata["scalars"] = mesh.coordinates[:, 2]
printc("is_closed :", mesh.is_closed())
printc("is_manifold:", mesh.is_manifold())
printc("euler_char :", mesh.euler_characteristic())
printc("genus :", mesh.genus())
reeb = mesh.to_reeb_graph()
ids = reeb[0].pointdata["Vertex Ids"]
pts = Points(mesh.coordinates[ids], r=10)
show([[mesh, pts], reeb], N=2, sharecam=False)
Source code in vedo/mesh/core.py
321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 | |
triangulate(verts=True, lines=True)
Converts mesh polygons into triangles.
If the input mesh is only made of 2D lines (no faces) the output will be a triangulation that fills the internal area. The contours may be concave, and may even contain holes, i.e. a contour may contain an internal contour winding in the opposite direction to indicate that it is a hole.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
verts
|
bool
|
if True, break input vertex cells into individual vertex cells (one point per cell). If False, the input vertex cells will be ignored. |
True
|
lines
|
bool
|
if True, break input polylines into line segments. If False, input lines will be ignored and the output will have no lines. |
True
|
Source code in vedo/mesh/core.py
677 678 679 680 681 682 683 684 685 686 687 688 689 690 691 692 693 694 695 696 697 698 699 700 701 702 703 704 705 706 707 708 709 710 711 712 713 714 715 716 717 718 719 | |
metrics
MeshMetricsMixin
Source code in vedo/mesh/metrics.py
17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 | |
cell_normals
property
Retrieve face normals as a numpy array.
If need be normals are computed via compute_normals().
Check out also compute_normals(cells=True) and compute_normals_with_pca().
vertex_normals
property
Retrieve vertex normals as a numpy array.
If needed, normals are automatically computed via compute_normals().
Check out also compute_normals_with_pca().
area()
Compute the surface area of the mesh.
The mesh must be triangular for this to work.
To triangulate a mesh use mesh.triangulate().
Source code in vedo/mesh/metrics.py
99 100 101 102 103 104 105 106 107 108 109 110 | |
check_validity(tol=0)
Return a numpy array of possible problematic faces following this convention: - Valid = 0 - WrongNumberOfPoints = 1 - IntersectingEdges = 2 - IntersectingFaces = 4 - NoncontiguousEdges = 8 - Nonconvex = 10 - OrientedIncorrectly = 20
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
tol
|
float
|
value is used as an epsilon for floating point equality checks throughout the cell checking process. |
0
|
Source code in vedo/mesh/metrics.py
321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 | |
compute_cell_vertex_count()
Add to this mesh a cell data array containing the nr of vertices that a polygonal face has.
Source code in vedo/mesh/metrics.py
239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 | |
compute_curvature(method=0)
Add scalars to Mesh that contains the curvature calculated in three different ways.
Variable method can be:
- 0 = gaussian
- 1 = mean curvature
- 2 = max curvature
- 3 = min curvature
Examples:
from vedo import Torus
Torus().compute_curvature().add_scalarbar().show().close()
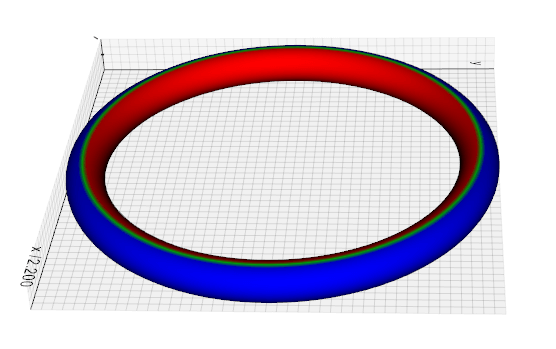
Source code in vedo/mesh/metrics.py
345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 | |
compute_elevation(low=(0, 0, 0), high=(0, 0, 1), vrange=(0, 1))
Add to Mesh a scalar array that contains distance along a specified direction.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
low
|
list
|
one end of the line (small scalar values) |
(0, 0, 0)
|
high
|
list
|
other end of the line (large scalar values) |
(0, 0, 1)
|
vrange
|
list
|
set the range of the scalar |
(0, 1)
|
Examples:
from vedo import Sphere
s = Sphere().compute_elevation(low=(0,0,0), high=(1,1,1))
s.add_scalarbar().show(axes=1).close()
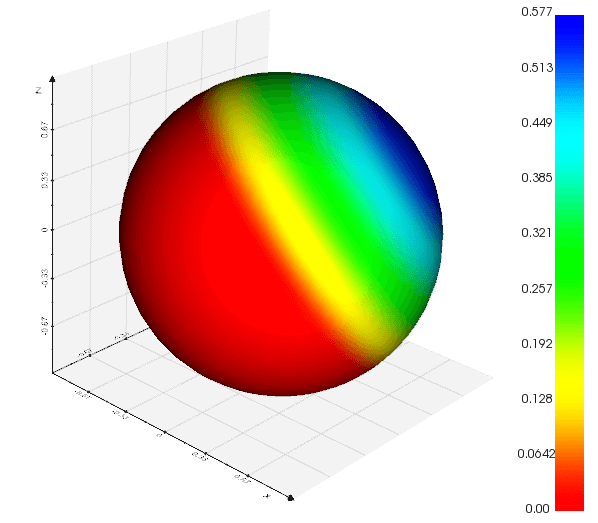
Source code in vedo/mesh/metrics.py
370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 | |
compute_normals(points=True, cells=True, feature_angle=None, consistency=True)
Compute cell and vertex normals for the mesh.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
points
|
bool
|
do the computation for the vertices too |
True
|
cells
|
bool
|
do the computation for the cells too |
True
|
feature_angle
|
float
|
specify the angle that defines a sharp edge. If the difference in angle across neighboring polygons is greater than this value, the shared edge is considered "sharp" and it is split. |
None
|
consistency
|
bool
|
turn on/off the enforcement of consistent polygon ordering. |
True
|
.. warning::
If feature_angle is set then the Mesh can be modified, and it
can have a different number of vertices from the original.
Note that the appearance of the mesh may change if the normals are computed,
as shading is automatically enabled when such information is present.
Use `mesh.flat()` to avoid smoothing effects.
Source code in vedo/mesh/metrics.py
44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 | |
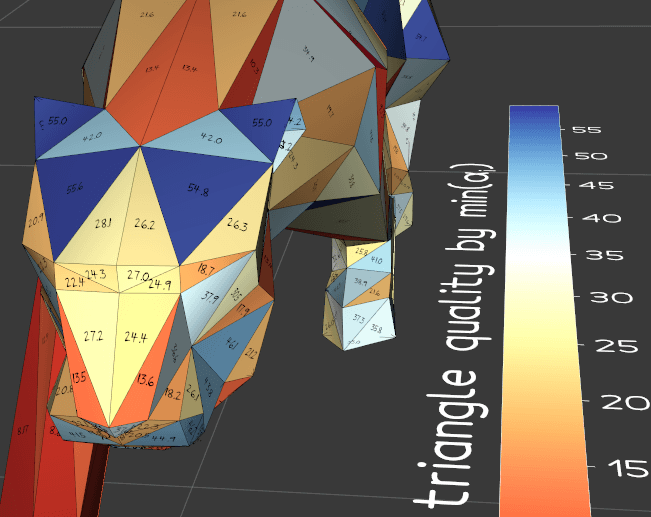
compute_quality(metric=6)
Calculate metrics of quality for the elements of a triangular mesh. This method adds to the mesh a cell array named "Quality". See class vtkMeshQuality.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
metric
|
int
|
type of available estimators are: - EDGE RATIO, 0 - ASPECT RATIO, 1 - RADIUS RATIO, 2 - ASPECT FROBENIUS, 3 - MED ASPECT FROBENIUS, 4 - MAX ASPECT FROBENIUS, 5 - MIN_ANGLE, 6 - COLLAPSE RATIO, 7 - MAX ANGLE, 8 - CONDITION, 9 - SCALED JACOBIAN, 10 - SHEAR, 11 - RELATIVE SIZE SQUARED, 12 - SHAPE, 13 - SHAPE AND SIZE, 14 - DISTORTION, 15 - MAX EDGE RATIO, 16 - SKEW, 17 - TAPER, 18 - VOLUME, 19 - STRETCH, 20 - DIAGONAL, 21 - DIMENSION, 22 - ODDY, 23 - SHEAR AND SIZE, 24 - JACOBIAN, 25 - WARPAGE, 26 - ASPECT GAMMA, 27 - AREA, 28 - ASPECT BETA, 29 |
6
|
Examples:

Source code in vedo/mesh/metrics.py
256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 | |
count_vertices()
Count the number of vertices each cell has and return it as a numpy array
Source code in vedo/mesh/metrics.py
312 313 314 315 316 317 318 319 | |
euler_characteristic()
Compute the Euler characteristic of the mesh. The Euler characteristic is a topological invariant for surfaces.
Source code in vedo/mesh/metrics.py
224 225 226 227 228 229 | |
genus()
Compute the genus of the mesh. The genus is a topological invariant for surfaces.
Source code in vedo/mesh/metrics.py
231 232 233 234 235 236 237 | |
is_closed()
Return True if the mesh is watertight.
Note that if the mesh contains coincident points the result may be flase.
Use in this case mesh.clean() to merge coincident points.
Source code in vedo/mesh/metrics.py
112 113 114 115 116 117 118 119 120 121 122 123 124 125 | |
is_manifold()
Return True if the mesh is manifold.
Source code in vedo/mesh/metrics.py
127 128 129 130 131 132 133 134 135 136 | |
non_manifold_faces(remove=True, tol='auto')
Detect and (try to) remove non-manifold faces of a triangular mesh:
- set `remove` to `False` to mark cells without removing them.
- set `tol=0` for zero-tolerance, the result will be manifold but with holes.
- set `tol>0` to cut off non-manifold faces, and try to recover the good ones.
- set `tol="auto"` to make an automatic choice of the tolerance.
Source code in vedo/mesh/metrics.py
138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 | |
volume()
Compute the volume occupied by mesh.
The mesh must be triangular for this to work.
To triangulate a mesh use mesh.triangulate().
Source code in vedo/mesh/metrics.py
86 87 88 89 90 91 92 93 94 95 96 97 | |